A cyclic peptide toolkit reveals mechanistic principles of peptidylarginine deiminase IV regulation.
Bertran, M.T., Walmsley, R., Cummings, T., Aramburu, I.V., Benton, D.J., Mora Molina, R., Assalaarachchi, J., Chasampalioti, M., Swanton, T., Joshi, D., Federico, S., Okkenhaug, H., Yu, L., Oxley, D., Walker, S., Papayannopoulos, V., Suga, H., Christophorou, M.A., Walport, L.J.(2024) Nat Commun 15: 9746-9746
- PubMed: 39528459 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-53554-1
- Primary Citation Related Structures:
8R8U, 8R8V - PubMed Abstract:
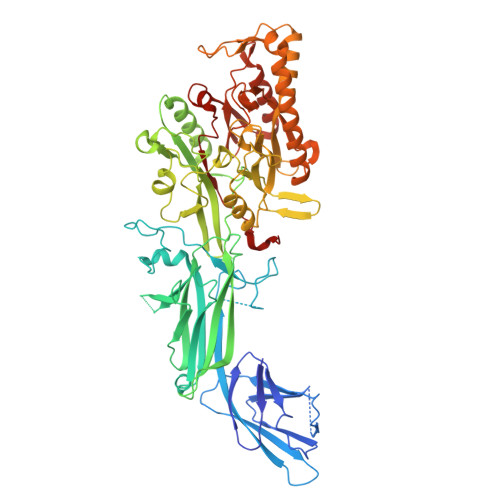
Peptidylarginine deiminase IV (PADI4, PAD4) deregulation promotes the development of autoimmunity, cancer, atherosclerosis and age-related tissue fibrosis. PADI4 additionally mediates immune responses and cellular reprogramming, although the full extent of its physiological roles is unexplored. Despite detailed molecular knowledge of PADI4 activation in vitro, we lack understanding of its regulation within cells, largely due to a lack of appropriate systems and tools. Here, we develop and apply a set of potent and selective PADI4 modulators. Using the mRNA-display-based RaPID system, we screen >10 12 cyclic peptides for high-affinity, conformation-selective binders. We report PADI4_3, a cell-active inhibitor specific for the active conformation of PADI4; PADI4_7, an inert binder, which we functionalise for the isolation and study of cellular PADI4; and PADI4_11, a cell-active PADI4 activator. Structural studies with PADI4_11 reveal an allosteric binding mode that may reflect the mechanism that promotes cellular PADI4 activation. This work contributes to our understanding of PADI4 regulation and provides a toolkit for the study and modulation of PADI4 across (patho)physiological contexts.
- Protein-Protein Interaction Laboratory, The Francis Crick Institute, London, NW1 1AT, UK.
Organizational Affiliation: