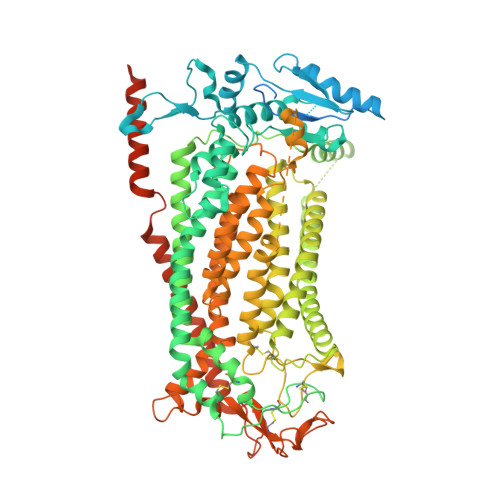
Mechanistic basis of ligand efficacy in the calcium-activated chloride channel TMEM16A.
Lam, A.K., Dutzler, R.(2023) EMBO J 42: e115030-e115030
- PubMed: 37984335 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2023115030
- Primary Citation Related Structures:
8QZC - PubMed Abstract:
Agonist binding in ligand-gated ion channels is coupled to structural rearrangements around the binding site, followed by the opening of the channel pore. In this process, agonist efficacy describes the equilibrium between open and closed conformations in a fully ligand-bound state. Calcium-activated chloride channels in the TMEM16 family are important sensors of intracellular calcium signals and are targets for pharmacological modulators, yet a mechanistic understanding of agonist efficacy has remained elusive. Using a combination of cryo-electron microscopy, electrophysiology, and autocorrelation analysis, we now show that agonist efficacy in the ligand-gated channel TMEM16A is dictated by the conformation of the pore-lining helix α6 around the Ca 2+ -binding site. The closure of the binding site, which involves the formation of a π-helix below a hinge region in α6, appears to be coupled to the opening of the inner pore gate, thereby governing the channel's open probability and conductance. Our results provide a mechanism for agonist binding and efficacy and a structural basis for the design of potentiators and partial agonists in the TMEM16 family.
- Department of Biochemistry, University of Zurich, Zurich, Switzerland.
Organizational Affiliation: