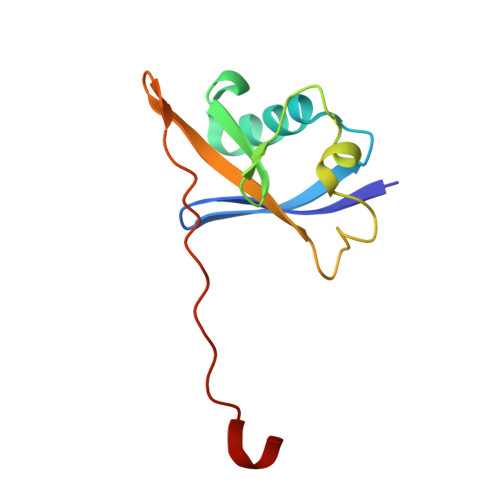
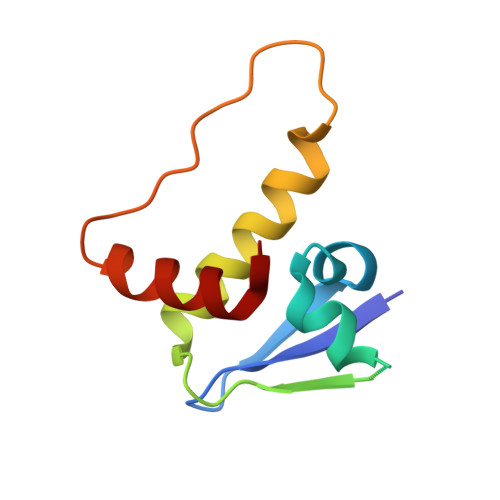
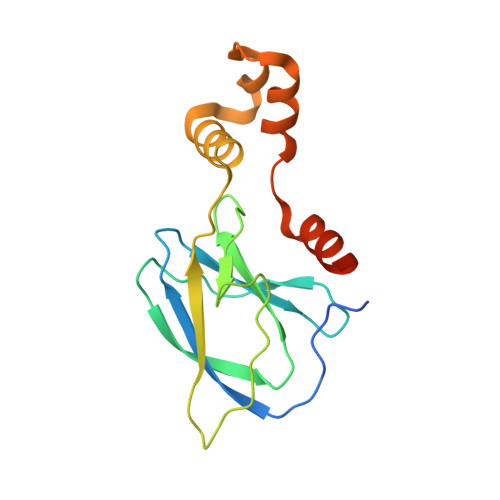
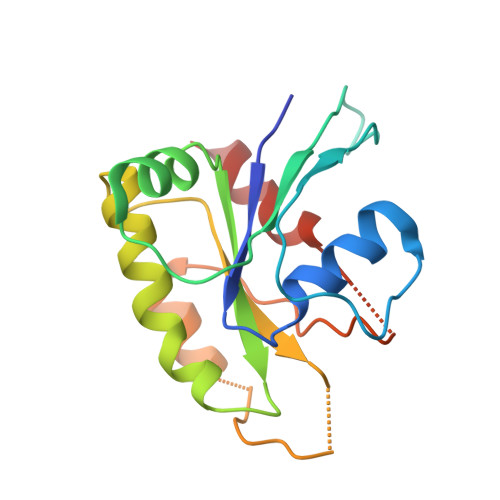
Targeting cancer with small-molecule pan-KRAS degraders.
Popow, J., Farnaby, W., Gollner, A., Kofink, C., Fischer, G., Wurm, M., Zollman, D., Wijaya, A., Mischerikow, N., Hasenoehrl, C., Prokofeva, P., Arnhof, H., Arce-Solano, S., Bell, S., Boeck, G., Diers, E., Frost, A.B., Goodwin-Tindall, J., Karolyi-Oezguer, J., Khan, S., Klawatsch, T., Koegl, M., Kousek, R., Kratochvil, B., Kropatsch, K., Lauber, A.A., McLennan, R., Olt, S., Peter, D., Petermann, O., Roessler, V., Stolt-Bergner, P., Strack, P., Strauss, E., Trainor, N., Vetma, V., Whitworth, C., Zhong, S., Quant, J., Weinstabl, H., Kuster, B., Ettmayer, P., Ciulli, A.(2024) Science 385: 1338-1347
- PubMed: 39298590 Search on PubMed
- DOI: https://doi.org/10.1126/science.adm8684
- Primary Citation Related Structures:
8QU8, 8QUG, 8QVU, 8QW6, 8QW7 - PubMed Abstract:
Mutations in the Kirsten rat sarcoma viral oncogene homolog (KRAS) protein are highly prevalent in cancer. However, small-molecule concepts that address oncogenic KRAS alleles remain elusive beyond replacing glycine at position 12 with cysteine (G12C), which is clinically drugged through covalent inhibitors. Guided by biophysical and structural studies of ternary complexes, we designed a heterobifunctional small molecule that potently degrades 13 out of 17 of the most prevalent oncogenic KRAS alleles. Compared with inhibition, KRAS degradation results in more profound and sustained pathway modulation across a broad range of KRAS mutant cell lines, killing cancer cells while sparing models without genetic KRAS aberrations. Pharmacological degradation of oncogenic KRAS was tolerated and led to tumor regression in vivo. Together, these findings unveil a new path toward addressing KRAS-driven cancers with small-molecule degraders.
- Boehringer Ingelheim RCV GmbH & Co KG, 1221 Vienna, Austria.
Organizational Affiliation: