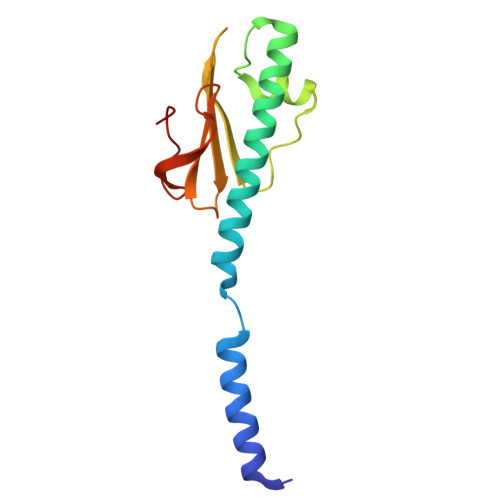
Structural insights into the Thermus thermophilus type IV pilus machinery assembling two distinct pili.
Neuhaus, A., McLaren, M., Isupov, M.N., Gaines, M., Buzzard, E., Sikora, M., Hanus, C., Daum, B., Averhoff, B., Gold, V.A.M.(2026) Commun Biol 9
- PubMed: 41917195 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-026-09762-0
- Primary Citation Related Structures:
8QQD, 8QQJ - PubMed Abstract:
Type IV pili are long, filamentous structures that extend from bacterial cell surfaces, enabling cells to respond to changing environments and facilitating genome plasticity. Thermus thermophilus HB27 produces two different type IV pili, each exhibiting distinct structural and functional properties. Here, we combine cryo-electron tomography, mutagenesis, and AlphaFold predictions to generate hypothetical in situ models of the T. thermophilus type IV pilus assembly machinery. Using single-particle cryo-electron microscopy, we determine structures of both filament types, enabling modelling of their surface glycans. Molecular dynamics simulations further reveal the flexibility of these glycans on extrusion. Integration of the filament structures with our hypothetical model of the assembly machinery offers a framework for further dissecting T4P architecture and biogenesis.
- Living Systems Institute, University of Exeter, Stocker Road, Exeter, UK.
Organizational Affiliation: