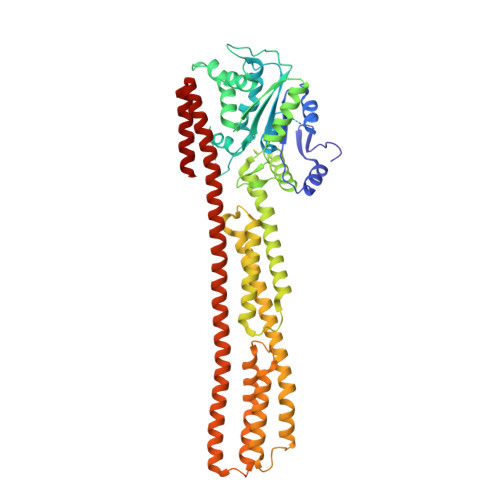
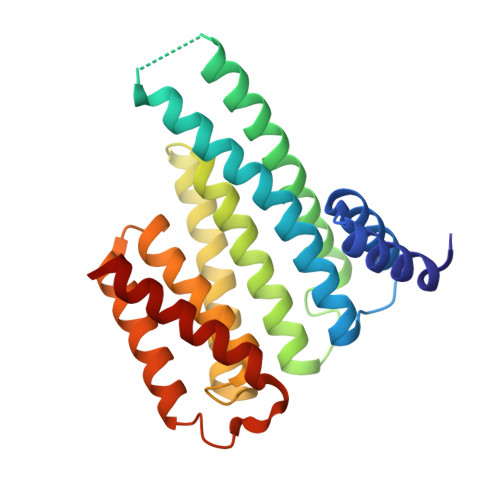
PIM1 controls GBP1 activity to limit self-damage and to guard against pathogen infection.
Fisch, D., Pfleiderer, M.M., Anastasakou, E., Mackie, G.M., Wendt, F., Liu, X., Clough, B., Lara-Reyna, S., Encheva, V., Snijders, A.P., Bando, H., Yamamoto, M., Beggs, A.D., Mercer, J., Shenoy, A.R., Wollscheid, B., Maslowski, K.M., Galej, W.P., Frickel, E.M.(2023) Science 382: eadg2253-eadg2253
- PubMed: 37797010 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.adg2253
- Primary Citation Related Structures:
8Q4L - PubMed Abstract:
Disruption of cellular activities by pathogen virulence factors can trigger innate immune responses. Interferon-γ (IFN-γ)-inducible antimicrobial factors, such as the guanylate binding proteins (GBPs), promote cell-intrinsic defense by attacking intracellular pathogens and by inducing programmed cell death. Working in human macrophages, we discovered that GBP1 expression in the absence of IFN-γ killed the cells and induced Golgi fragmentation. IFN-γ exposure improved macrophage survival through the activity of the kinase PIM1. PIM1 phosphorylated GBP1, leading to its sequestration by 14-3-3σ, which thereby prevented GBP1 membrane association. During Toxoplasma gondii infection, the virulence protein TgIST interfered with IFN-γ signaling and depleted PIM1, thereby increasing GBP1 activity. Although infected cells can restrain pathogens in a GBP1-dependent manner, this mechanism can protect uninfected bystander cells. Thus, PIM1 can provide a bait for pathogen virulence factors, guarding the integrity of IFN-γ signaling.
- Host-Toxoplasma Interaction Laboratory, The Francis Crick Institute, London, UK.
Organizational Affiliation: