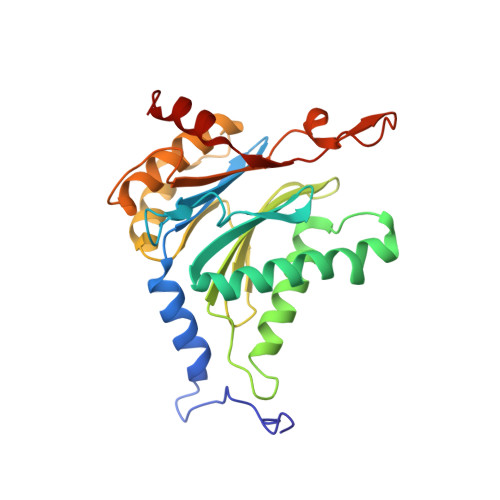
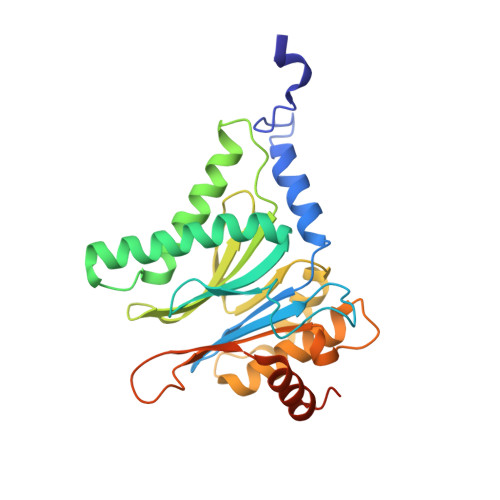
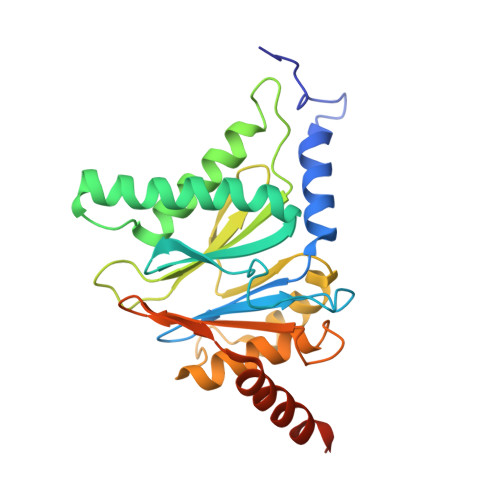
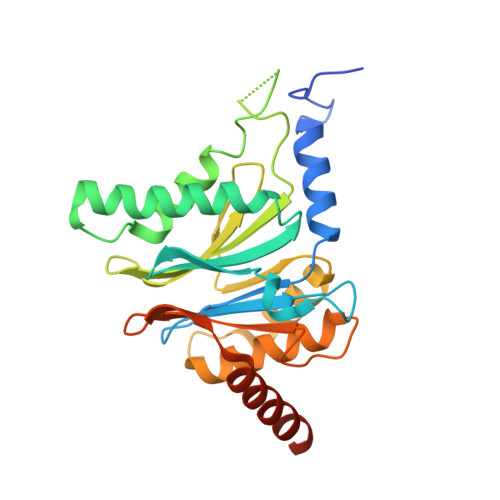
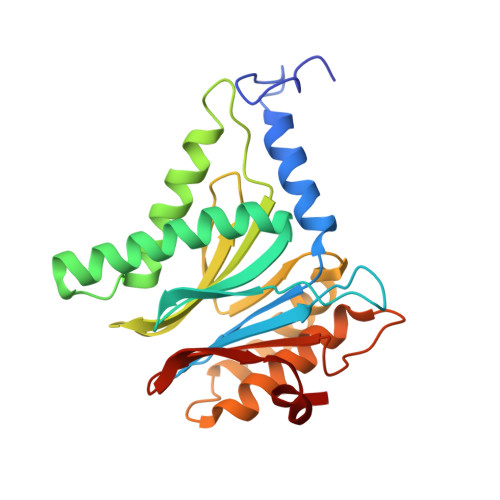
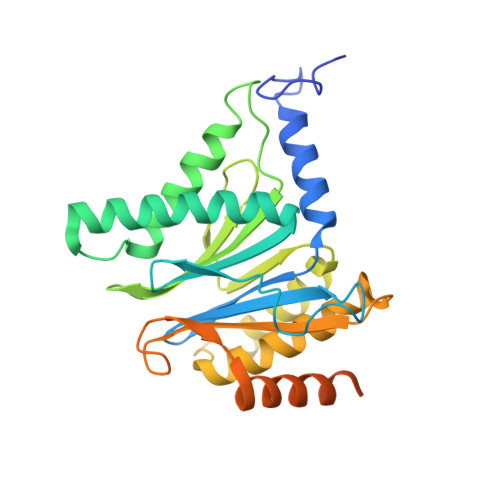
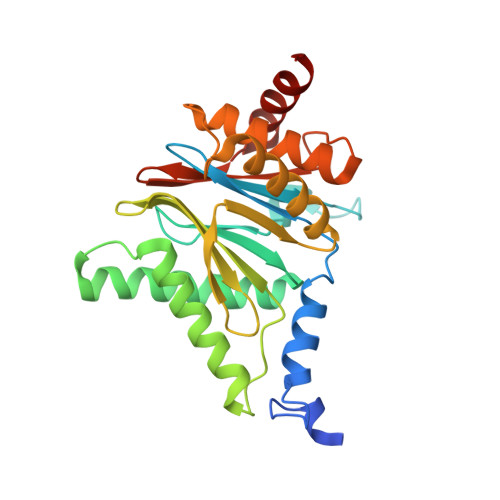
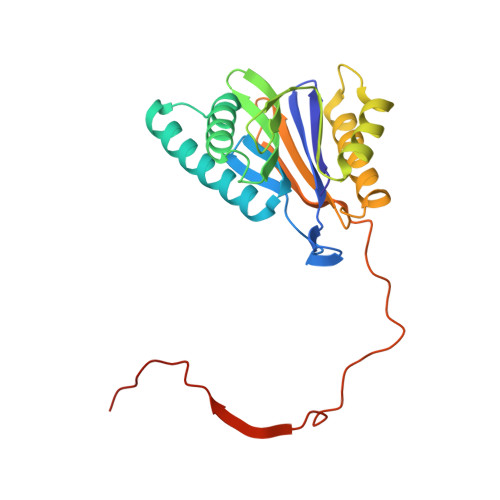
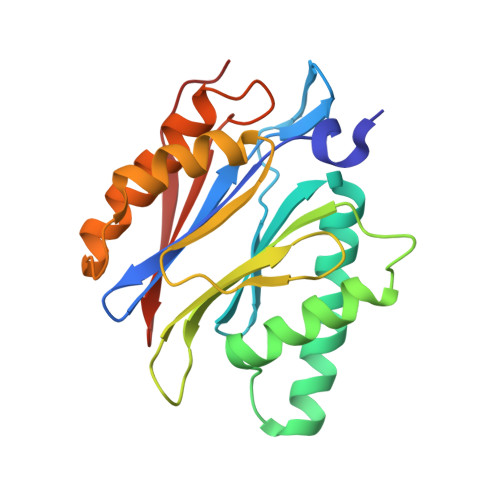
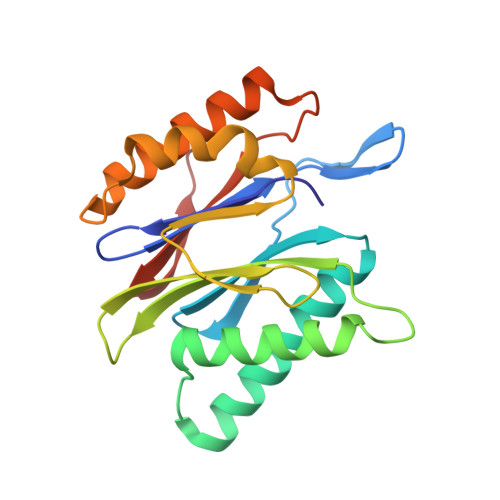
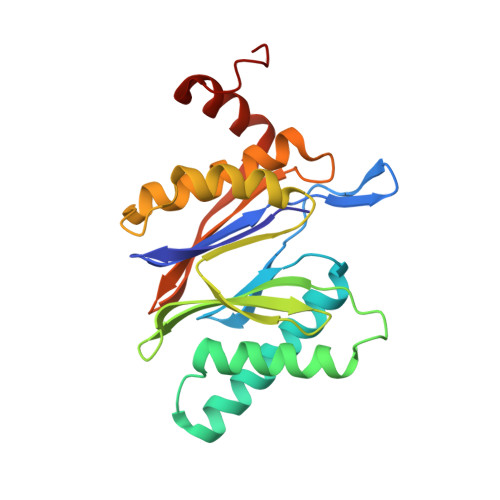
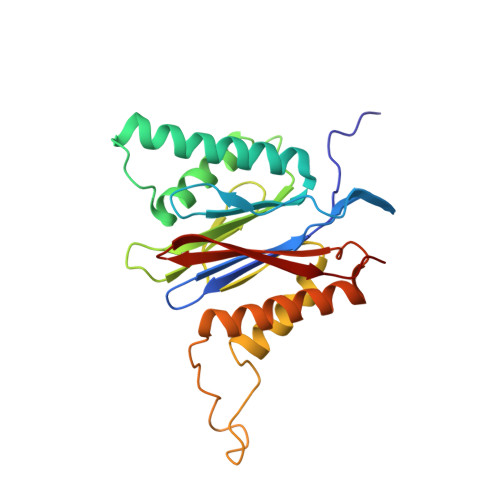
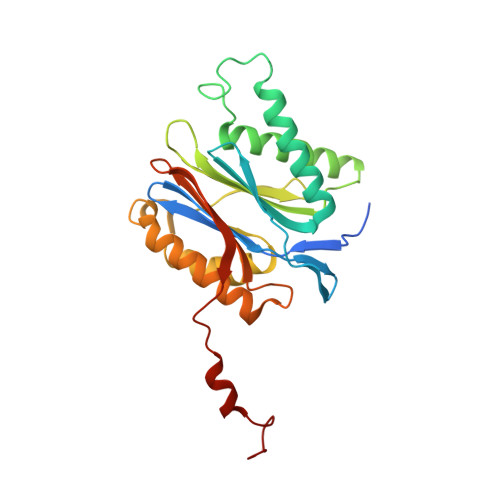
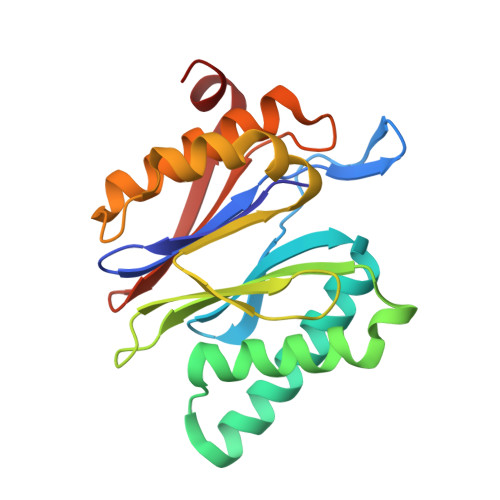
Structure of Staphylococcus aureus ClpP Bound to the Covalent Active-Site Inhibitor Cystargolide A.
Illigmann, A., Vielberg, M.T., Lakemeyer, M., Wolf, F., Dema, T., Stange, P., Kuttenlochner, W., Liebhart, E., Kulik, A., Staudt, N.D., Malik, I., Grond, S., Sieber, S.A., Kaysser, L., Groll, M., Brotz-Oesterhelt, H.(2024) Angew Chem Int Ed Engl 63: e202314028-e202314028
- PubMed: 38029352 Search on PubMed
- DOI: https://doi.org/10.1002/anie.202314028
- Primary Citation Related Structures:
8OLL, 8OLR, 8R03, 8R04, 8R05 - PubMed Abstract:
The caseinolytic protease is a highly conserved serine protease, crucial to prokaryotic and eukaryotic protein homeostasis, and a promising antibacterial and anticancer drug target. Herein, we describe the potent cystargolides as the first natural β-lactone inhibitors of the proteolytic core ClpP. Based on the discovery of two clpP genes next to the cystargolide biosynthetic gene cluster in Kitasatospora cystarginea, we explored ClpP as a potential cystargolide target. We show the inhibition of Staphylococcus aureus ClpP by cystargolide A and B by different biochemical methods in vitro. Synthesis of semisynthetic derivatives and probes with improved cell penetration allowed us to confirm ClpP as a specific target in S. aureus cells and to demonstrate the anti-virulence activity of this natural product class. Crystal structures show cystargolide A covalently bound to all 14 active sites of ClpP from S. aureus, Aquifex aeolicus, and Photorhabdus laumondii, and reveal the molecular mechanism of ClpP inhibition by β-lactones, the predominant class of ClpP inhibitors.
- Department of Microbial Bioactive Compounds, Interfaculty Institute of Microbiology and Infection Medicine, University of Tübingen, Auf der Morgenstelle 28, 72076, Tübingen, Germany.
Organizational Affiliation: