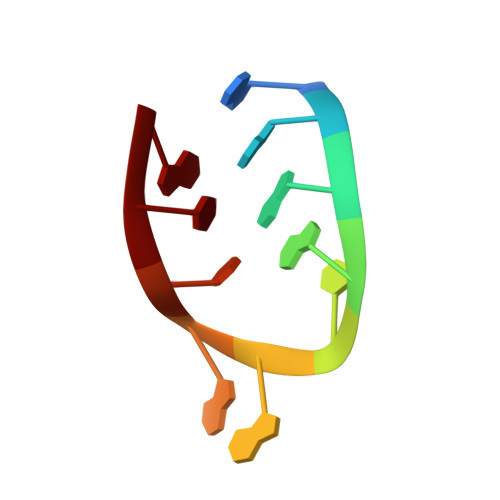Structural basis for the activation of plant bunyavirus replication machinery and its dual-targeted inhibition by ribavirin.
Li, J., Cao, L., Zhao, Y., Shen, J., Wang, L., Feng, M., Zhu, M., Ye, Y., Kormelink, R., Tao, X., Wang, X.(2025) Nat Plants 11: 518-530
- PubMed: 40044941 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41477-025-01940-y
- Primary Citation Related Structures:
8KI6, 8KI7, 8KI8, 8KI9, 8KIA, 9J8V - PubMed Abstract:
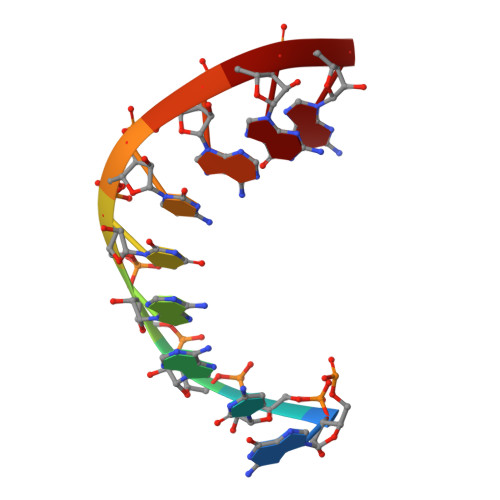
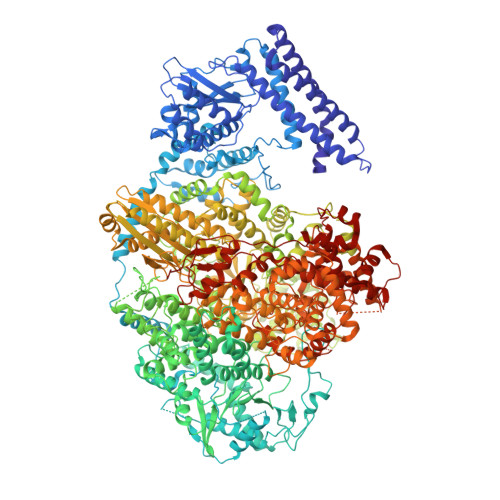
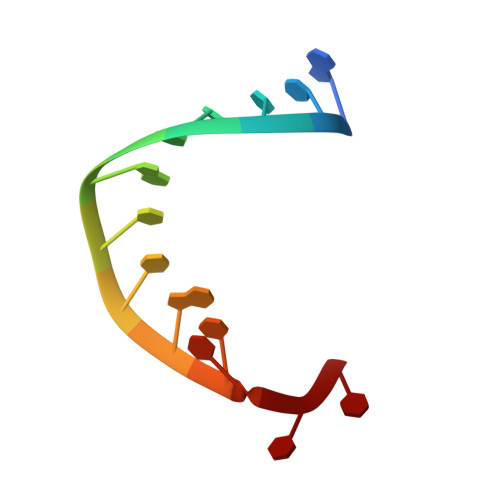
Despite the discovery of plant viruses as a new class of pathogens over a century ago, the structure of plant virus replication machinery and antiviral pesticide remains lacking. Here we report five cryogenic electron microscopy structures of a ~330-kDa RNA-dependent RNA polymerase (RdRp) from a devastating plant bunyavirus, tomato spotted wilt orthotospovirus (TSWV), including the apo, viral-RNA-bound, base analogue ribavirin-bound and ribavirin-triphosphate-bound states. They reveal that a flexible loop of RdRp's motif F functions as 'sensor' to perceive viral RNA and further acts as an 'adaptor' to promote the formation of a complete catalytic centre. A ten-base RNA 'hook' structure is sufficient to trigger major conformational changes and activate RdRp. Chemical screening showed that ribavirin is effective against TSWV, and structural data revealed that ribavirin disrupts both hook-binding and catalytic core formation, locking polymerase in its inactive state. This work provides structural insights into the mechanisms of plant bunyavirus RdRp activation and its dual-targeted site inhibition, facilitating the development of pesticides against plant viruses.
- State Key Laboratory of Agricultural and Forestry Biosecurity, Nanjing Agricultural University, Nanjing, China.
Organizational Affiliation: