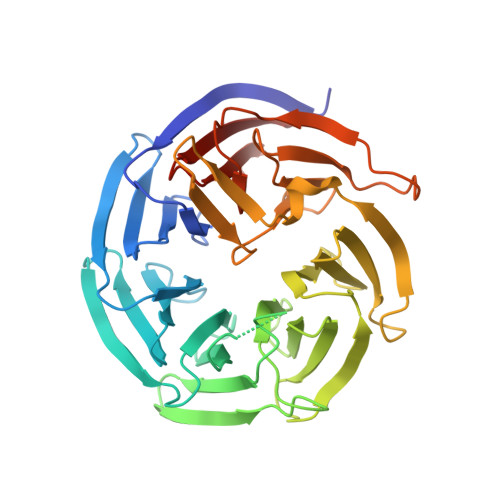
Insights into the distinct membrane targeting mechanisms of WDR91 family proteins.
Ma, X., Li, J., Liu, N., Banerjee, S., Hu, X., Wang, X., Dong, J., Liu, K., Yang, C., Dong, Z.(2024) Structure 32: 2287-2300.e4
- PubMed: 39426373 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2024.09.023
- Primary Citation Related Structures:
8KB8, 8KB9 - PubMed Abstract:
WDR91 and SORF1, members of the WD repeat-containing protein 91 family, control phosphoinositide conversion by inhibiting phosphatidylinositol 3-kinase activity on endosomes, which promotes endosome maturation. Here, we report the crystal structure of the human WDR91 WD40 domain complexed with Rab7 that has an unusual interface at the C-terminus of the Rab7 switch II region. WDR91 is highly selective for Rab7 among the tested GTPases. A LIS1 homology (LisH) motif within the WDR91 N-terminal domain (NTD) mediates self-association and may contribute partly to the augmented interaction between full-length WDR91 and Rab7. Both the Rab7 binding site and the LisH motif are indispensable for WDR91 function in endocytic trafficking. For the WDR91 orthologue SORF1 lacking the C-terminal WD40 domain, a C-terminal amphipathic helix (AH) mediates strong interactions with liposomes containing acidic lipids. During evolution the human WDR91 ancestor gene might have acquired a WD40 domain to replace the AH for endosomal membrane targeting.
- China-US (Henan) Hormel Cancer Institute, Zhengzhou, Henan 450003, China.
Organizational Affiliation: