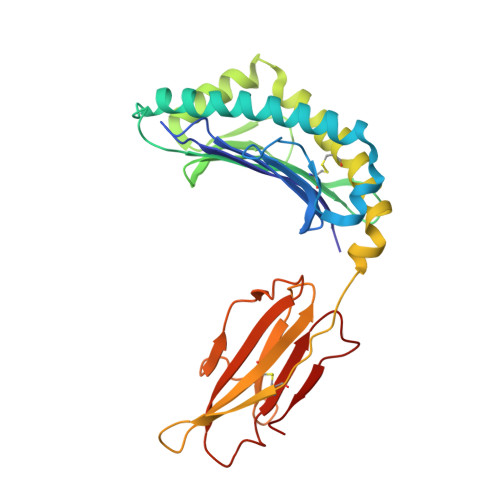
Structural analyses of Cryptosporidium parvum epitopes reveal a novel scheme of decapeptide binding to H-2K b.
Wang, Y., Chang, Y., Yin, F., Kang, C., Meng, Y., Xu, F., Liu, Y., Zhang, Y., Wu, C., Fan, S., Zhao, J.(2025) J Struct Biol 217: 108168-108168
- PubMed: 39809366 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2025.108168
- Primary Citation Related Structures:
8JHW - PubMed Abstract:
Cryptosporidium has gained much attention as a major cause of diarrhea worldwide. Here, we present the first structure of H-2K b complexed with a decapeptide from Cryptosporidium parvum Gp40/15 protein (Gp40/15-VTF10). In contrast to all published structures, the aromatic residue P3-Phe of Gp40/15-VTF10 is anchored in pocket C rather than the canonical Y/F at P5 or P6 reported for octapeptides and nonapeptides. The results of in vitro refolding assays and circular dichroism experiments showed that the side chains of P3 and P5 play key roles in Gp40/15-VTF10 peptide binding. However, functional analysis of decapeptide epitopes revealed that the Gp40/15-VTF10 peptide did not elicit a strong CD8 + T immune response, whereas the decapeptide epitope MEDLE2-INF10 induced a significant CD8 + T-cell response in peptide-immunized C57BL/6 mice. Using a model structure of H-2K b -INF10 complex, we found that the antigenic decapeptide INF10 exhibits a completely different conformation, with the aromatic anchors P3F and P7F docked into the D and C pockets, respectively, while similar peptide conformation and hydrogen bond interactions between the peptide and major histocompatibility complex were found in the resolved H-2K b -SVF9 complex. As the H-2K b molecule predominantly prefers octapeptides with a strong anchor of P5 Y/F (or P6 Y/F for nonapeptides) binding to the C pocket, we propose that P7 Y/F in the C pocket may represent a novel binding mode for decapeptides. The results should increase the accuracy of T-cell epitope prediction and support the development of T-cell epitope vaccines against cryptosporidiosis.
- State Key Laboratory of Agricultural Microbiology, College of Veterinary Medicine, Huazhong Agricultural University, Wuhan, PR China; College of Life Science and Agronomy, Zhoukou Normal University, Zhoukou, PR China.
Organizational Affiliation: