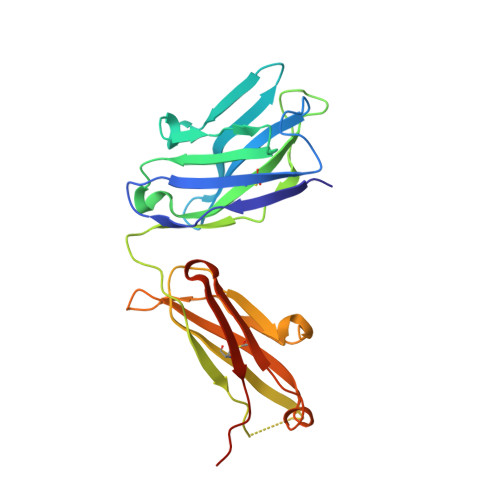
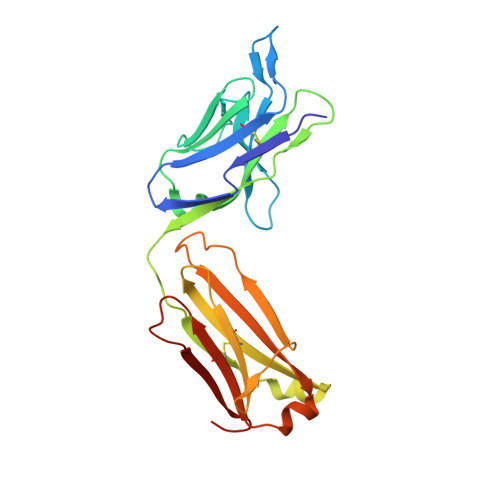
Structural insights into the unique pH-responsive characteristics of the anti-TIGIT therapeutic antibody Ociperlimab.
Sun, J., Zhang, X., Xue, L., Cheng, L., Zhang, J., Chen, X., Shen, Z., Li, K., Wang, L., Huang, C., Song, J.(2024) Structure 32: 550
- PubMed: 38460520 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2024.02.009
- Primary Citation Related Structures:
8JEL, 8JEN, 8JEO, 8JEP, 8JEQ - PubMed Abstract:
TIGIT is mainly expressed on T cells and is an inhibitory checkpoint receptor that binds to its ligand PVR in the tumor microenvironment. Anti-TIGIT monoclonal antibodies (mAbs) such as Ociperlimab and Tiragolumab block the TIGIT-PVR interaction and are in clinical development. However, the molecular blockade mechanism of these mAbs remains elusive. Here, we report the crystal structures of TIGIT in complex with Ociperlimab_Fab and Tiragolumab_Fab revealing that both mAbs bind TIGIT with a large steric clash with PVR. Furthermore, several critical epitopic residues are identified. Interestingly, the binding affinity of Ociperlimab toward TIGIT increases approximately 17-fold when lowering the pH from 7.4 to 6.0. Our structure shows a strong electrostatic interaction between ASP103 HCDR3 and HIS76 TIGIT explaining the pH-responsive mechanism of Ociperlimab. In contrast, Tiragolumab does not show an acidic pH-dependent binding enhancement. Our results provide valuable information that could help to improve the efficacy of therapeutic antibodies for cancer treatment.
- Department of Biologics, BeiGene (Beijing) Co., Ltd, Beijing, China.
Organizational Affiliation: