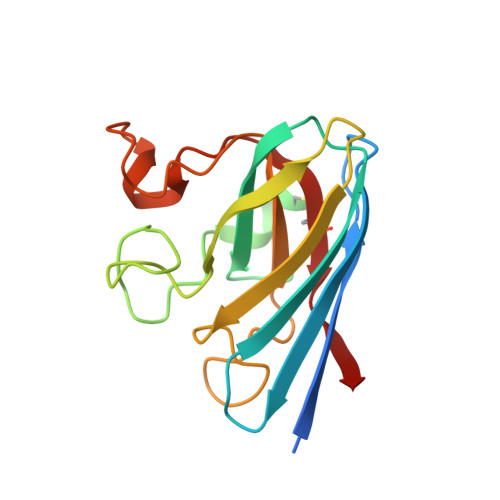
Direct relationship between dimeric form and activity in the acidic copper-zinc superoxide dismutase from lemon.
Utami, R.A., Yoshida, H., Kartadinata, L.H., Abdillah, V.A., Faratilla, C.R., Retnoningrum, D.S., Ismaya, W.T.(2023) Acta Crystallogr F Struct Biol Commun 79: 301-307
- PubMed: 38108885 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X23010646
- Primary Citation Related Structures:
8JD8 - PubMed Abstract:
The copper-zinc superoxide dismutase (CuZnSOD) from lemon (SOD_CL) is active in an acidic environment and resists proteolytic degradation. The enzyme occurs as a dimer, which has an indirect effect on the enzyme activity as the monomer retains only ∼35% of the activity. Here, the crystal structure of SOD_CL at 1.86 Å resolution is reported that may explain this peculiarity. The crystal belonged to space group P2 1 , with unit-cell parameters a = 61.11, b = 74.55, c = 61.69 Å, β = 106.86°, and contained four molecules in the asymmetric unit. The overall structure of SOD_CL resembles that of CuZnSOD from plants. The structure of SOD_CL shows a unique arrangement of surface loop IV that connects the dimer interface and the active site, which is located away from the dimer-interface region. This arrangement allows direct interaction between the residues residing in the dimer interface and those in the active site. The arrangement also includes Leu62 and Gln164, which are conserved in cytoplasmic CuZnSOD. This supports the classification of SOD_CL as a cytoplasmic CuZnSOD despite sharing the highest amino-acid sequence homology with CuZnSODs from spinach and tomato, which are chloroplastic.
- Laboratory of Pharmaceutical Biotechnology, Department of Pharmaceutics, School of Pharmacy, Bandung Institute of Technology, Jalan Ganesa No. 10, Bandung 40132, Indonesia.
Organizational Affiliation: