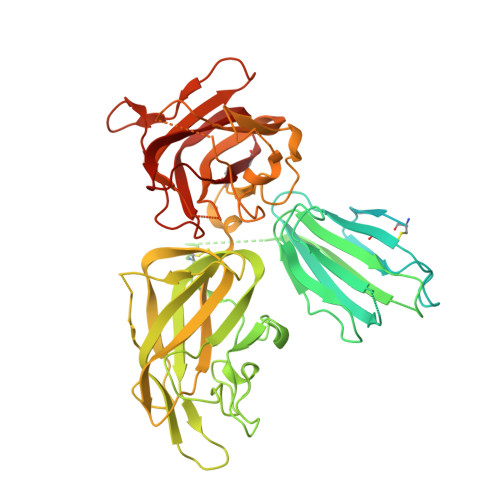
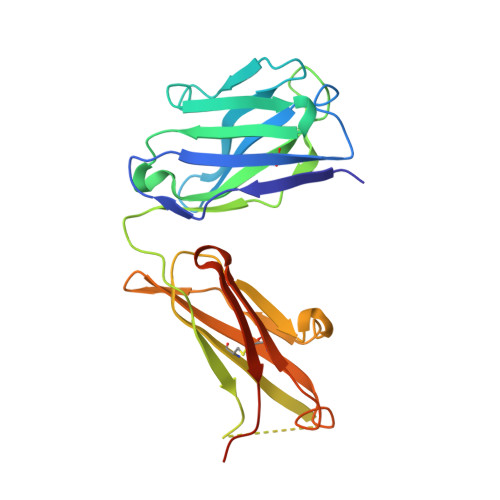
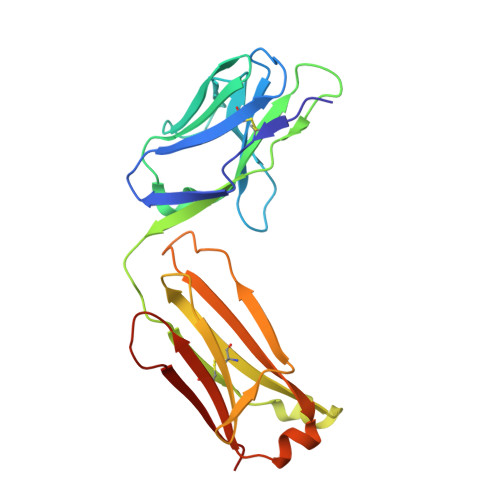
Inhibition of VEGF binding to neuropilin-2 enhances chemosensitivity and inhibits metastasis in triple-negative breast cancer.
Xu, Z., Goel, H.L., Burkart, C., Burman, L., Chong, Y.E., Barber, A.G., Geng, Y., Zhai, L., Wang, M., Kumar, A., Menefee, A., Polizzi, C., Eide, L., Rauch, K., Rahman, J., Hamel, K., Fogassy, Z., Klopp-Savino, S., Paz, S., Zhang, M., Cubitt, A., Nangle, L.A., Mercurio, A.M.(2023) Sci Transl Med 15: eadf1128-eadf1128
- PubMed: 37134152 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/scitranslmed.adf1128
- Primary Citation Related Structures:
8IVW, 8IVX - PubMed Abstract:
Although blocking the binding of vascular endothelial growth factor (VEGF) to neuropilin-2 (NRP2) on tumor cells is a potential strategy to treat aggressive carcinomas, a lack of effective reagents that can be used clinically has hampered this potential therapy. Here, we describe the generation of a fully humanized, high-affinity monoclonal antibody (aNRP2-10) that specifically inhibits the binding of VEGF to NRP2, conferring antitumor activity without causing toxicity. Using triple-negative breast cancer as a model, we demonstrated that aNRP2-10 could be used to isolate cancer stem cells (CSCs) from heterogeneous tumor populations and inhibit CSC function and epithelial-to-mesenchymal transition. aNRP2-10 sensitized cell lines, organoids, and xenografts to chemotherapy and inhibited metastasis by promoting the differentiation of CSCs to a state that is more responsive to chemotherapy and less prone to metastasis. These data provide justification for the initiation of clinical trials designed to improve the response of patients with aggressive tumors to chemotherapy using this monoclonal antibody.
- aTyr Pharma, San Diego, CA 92121, USA.
Organizational Affiliation: