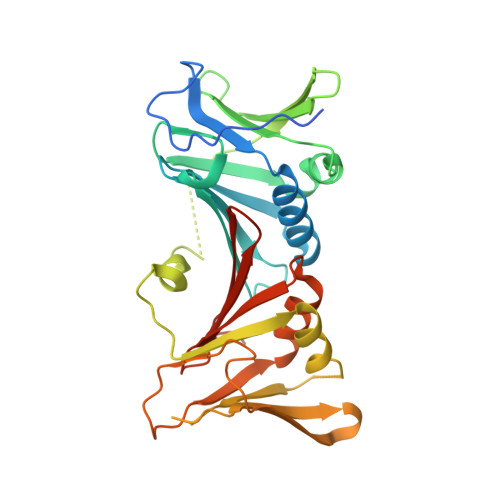
The E301R protein of African swine fever virus functions as a sliding clamp involved in viral genome replication.
Li, S., Ge, H., Li, Y., Zhang, K., Yu, S., Cao, H., Wang, Y., Deng, H., Li, J., Dai, J., Li, L.F., Luo, Y., Sun, Y., Geng, Z., Dong, Y., Zhang, H., Qiu, H.J.(2023) mBio 14: e0164523-e0164523
- PubMed: 37772878 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mbio.01645-23
- Primary Citation Related Structures:
8ITE - PubMed Abstract:
Sliding clamp is a highly conserved protein in the evolution of prokaryotic and eukaryotic cells. The sliding clamp is required for genomic replication as a critical co-factor of DNA polymerases. However, the sliding clamp analogs in viruses remain largely unknown. We found that the ASFV E301R protein (pE301R) exhibited a sliding clamp-like structure and similar functions during ASFV replication. Interestingly, pE301R is assembled into a unique ring-shaped homotetramer distinct from sliding clamps or proliferating cell nuclear antigens (PCNAs) from other species. Notably, the E301R gene is required for viral life cycle, but the pE301R function can be partially restored by the porcine PCNA. This study not only highlights the functional role of the ASFV pE301R as a viral sliding clamp analog, but also facilitates the dissection of the complex replication mechanism of ASFV, which provides novel clues for developing antivirals against ASF.
- State Key Laboratory for Animal Disease Control and Prevention, National African Swine Fever Para-Reference Laboratory, National High-Containment Facilities for Animal Disease Control and Prevention, Harbin Veterinary Research Institute, Chinese Academy of Agricultural Sciences , Harbin, China.
Organizational Affiliation: