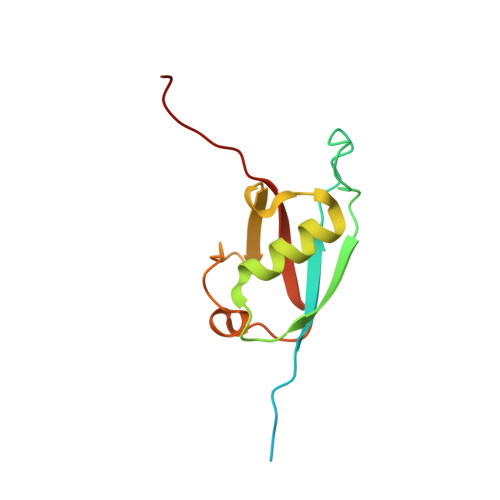
Structural insights into recognition of SL4, the UUCG stem-loop, of human U1 snRNA by the ubiquitin-like domain, including the C-terminal tail in the SF3A1 subunit of U2 snRNP.
Nameki, N., Terawaki, S.I., Takizawa, M., Kitamura, M., Muto, Y., Kuwasako, K.(2023) J Biochem 174: 203-216
- PubMed: 37094335 Search on PubMed
- DOI: https://doi.org/10.1093/jb/mvad033
- Primary Citation Related Structures:
8ID2 - PubMed Abstract:
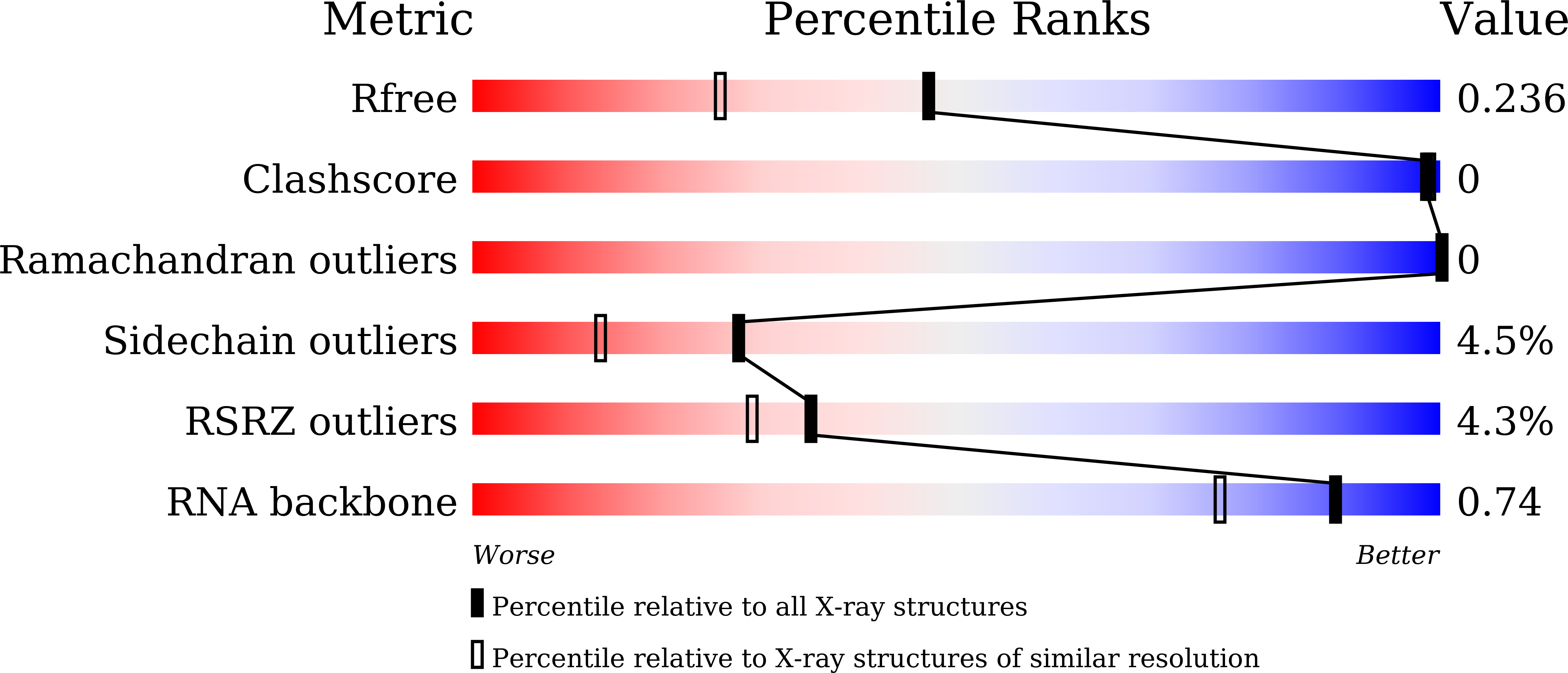
The pre-spliceosomal complex involves interactions between U1 and U2 snRNPs, where a ubiquitin-like domain (ULD) of SF3A1, a component of U2 snRNP, binds to the stem-loop 4 (SL4; the UUCG tetraloop) of U1 snRNA in U1 snRNP. Here, we reported the 1.80 Å crystal structure of human SF3A1 ULD (ULDSF3A1) complexed with SL4. The structural part of ULDSF3A1 (res. 704-785) adopts a typical β-grasp fold with a topology of β1-β2-α1-310a-β3-β4-310b-β5, closely resembling that of ubiquitin, except for the length and structure of the β1/β2 loop. A patch on the surface formed by three ULDSF3A1-specific residues, Lys756 (β3), Phe763 (β4) and Lys765 (following β4), contacts the canonical UUCG tetraloop structure. In contrast, the directly following C-terminal tail composed of 786KERGGRKK793 was essentially stretched. The main or side chains of all the residues interacted with the major groove of the stem helix; the RGG residues adopted a peculiar conformation for RNA recognition. These findings were confirmed by mutational studies using bio-layer interferometry. Collectively, a unique combination of the β-grasp fold and the C-terminal tail constituting ULDSF3A1 is required for the SL4-specific binding. This interaction mode also suggests that putative post-translational modifications, including ubiquitination in ULDSF3A1, directly inhibit SL4 binding.
- Division of Molecular Science, Graduate School of Science and Technology, Gunma University, 1-5-1 Tenjin-cho, Kiryu-shi, Gunma, 376-8515, Japan.
Organizational Affiliation: