Crystal structure of Co-type nitrile hydratase mutant L6T from Pseudomonas thermophila at 2.2 Angstroms resolution.
Cheng, Z.Y., Ma, D., Yin, M., Lai, Q.P., Peplowski, L., Han, L.C., Hou, X.D., Yin, D.J., Rao, Y.J., Zhou, Z.M.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cobalt-containing nitrile hydratase subunit alpha | 205 | Pseudonocardia thermophila | Mutation(s): 1 Gene Names: SAMN05443637_10360 EC: 4.2.1.84 | 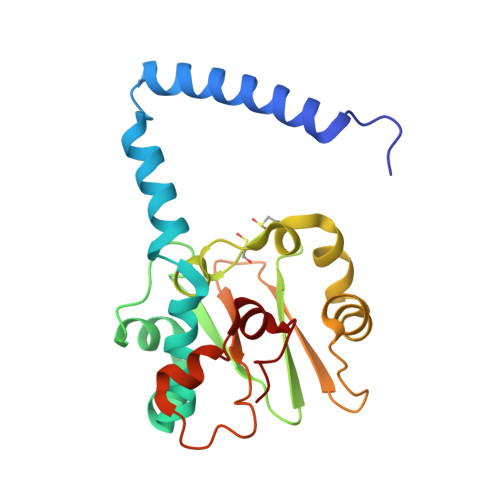 | |
UniProt | |||||
Find proteins for Q7SID2 (Pseudonocardia thermophila) Explore Q7SID2 Go to UniProtKB: Q7SID2 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q7SID2 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Cobalt-containing nitrile hydratase subunit beta | 233 | Pseudonocardia thermophila | Mutation(s): 0 EC: 4.2.1.84 | 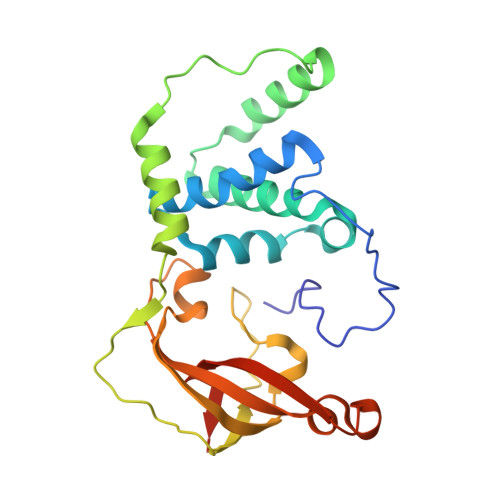 | |
UniProt | |||||
Find proteins for Q7SID3 (Pseudonocardia thermophila) Explore Q7SID3 Go to UniProtKB: Q7SID3 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q7SID3 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| CO (Subject of Investigation/LOI) Query on CO | C [auth A] | COBALT (II) ION Co XLJKHNWPARRRJB-UHFFFAOYSA-N |  | ||
| Modified Residues 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| CSD Query on CSD | A | L-PEPTIDE LINKING | C3 H7 N O4 S |  | CYS |
| CSO Query on CSO | A | L-PEPTIDE LINKING | C3 H7 N O3 S |  | CYS |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 65.64 | α = 90 |
| b = 65.64 | β = 90 |
| c = 184.07 | γ = 120 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| SAINT | data scaling |
| SAINT | data reduction |
| PHENIX | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China (NSFC) | China | -- |