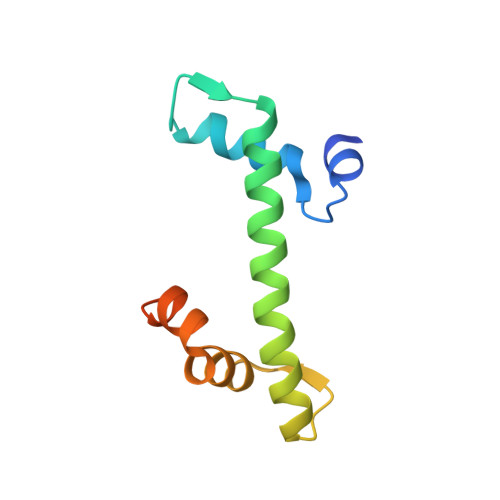
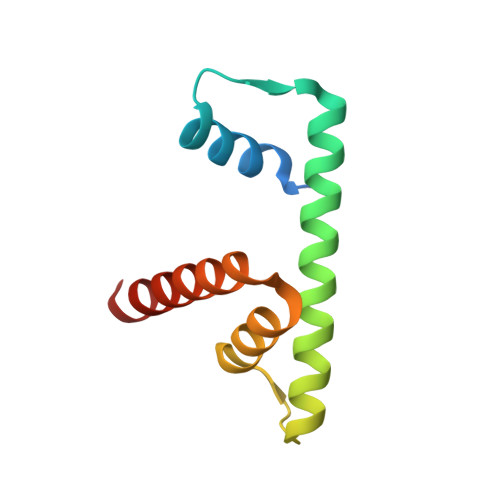
Structural basis for H2A-H2B recognitions by human Spt16.
Li, Y., Huang, H.(2023) Biochem Biophys Res Commun 651: 85-91
- PubMed: 36801613 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2023.02.016
- Primary Citation Related Structures:
8I17 - PubMed Abstract:
The human facilitates chromatin transcription (FACT) complex, consisting of Spt16 and SSRP1, is a versatile histone chaperone that can engage free H2A-H2B dimer and H3-H4 tetramer (or dimer), and partially unraveled nucleosome. The C-terminal domain of human Spt16 (hSpt16-CTD) is the decisive element for engaging H2A-H2B dimer and partially unraveled nucleosome. The molecular basis of the H2A-H2B dimer recognitions by hSpt16-CTD is not fully comprehended. Here, we present a high-resolution snapshot of the recognitions of the H2A-H2B dimer by hSpt16-CTD via an acidic intrinsically disordered (AID) segment, and reveal some distinct structural features of hSpt16-CTD as compared to the budding yeast Spt16-CTD.
- Key Laboratory of Molecular Design for Plant Cell Factory of Guangdong Higher Education Institutes, Department of Chemical Biology & Department of Biology, School of Life Sciences, Southern University of Science and Technology, Shenzhen, 518055, China.
Organizational Affiliation: