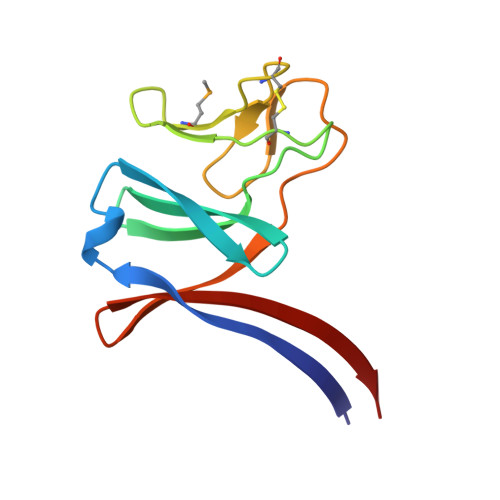
Soluble domains of cytochrome c-556 and Rieske iron-sulfur protein from Chlorobaculum tepidum: Crystal structures and interaction analysis.
Kishimoto, H., Azai, C., Yamamoto, T., Mutoh, R., Nakaniwa, T., Tanaka, H., Miyanoiri, Y., Kurisu, G., Oh-Oka, H.(2023) Curr Res Struct Biol 5: 100101-100101
- PubMed: 37180033 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.crstbi.2023.100101
- Primary Citation Related Structures:
8HN2, 8HN3 - PubMed Abstract:
In photosynthetic green sulfur bacteria, the electron transfer reaction from menaquinol:cytochrome c oxidoreductase to the P840 reaction center (RC) complex occurs directly without any involvement of soluble electron carrier protein(s). X-ray crystallography has determined the three-dimensional structures of the soluble domains of the CT0073 gene product and Rieske iron-sulfur protein (ISP). The former is a mono-heme cytochrome c with an α-absorption peak at 556 nm. The overall fold of the soluble domain of cytochrome c -556 (designated as cyt c -556 sol ) consists of four α-helices and is very similar to that of water-soluble cyt c -554 that independently functions as an electron donor to the P840 RC complex. However, the latter's remarkably long and flexible loop between the α3 and α4 helices seems to make it impossible to be a substitute for the former. The structure of the soluble domain of the Rieske ISP (Rieske sol protein) shows a typical β-sheets-dominated fold with a small cluster-binding and a large subdomain. The architecture of the Rieske sol protein is bilobal and belongs to those of b 6 f -type Rieske ISPs. Nuclear magnetic resonance (NMR) measurements revealed weak non-polar but specific interaction sites on Rieske sol protein when mixed with cyt c -556 sol . Therefore, menaquinol:cytochrome c oxidoreductase in green sulfur bacteria features a Rieske/cyt b complex tightly associated with membrane-anchored cyt c -556.
- Department of Biological Sciences, Graduate School of Science, Osaka University, Toyonaka, Osaka, 560-0043, Japan.
Organizational Affiliation: