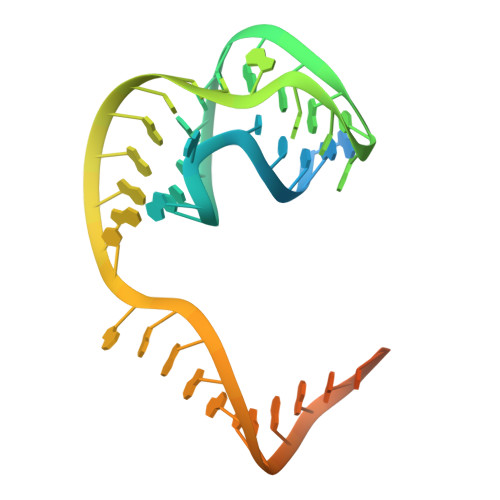
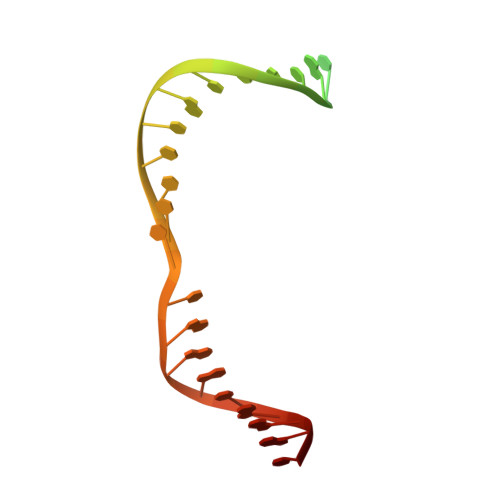
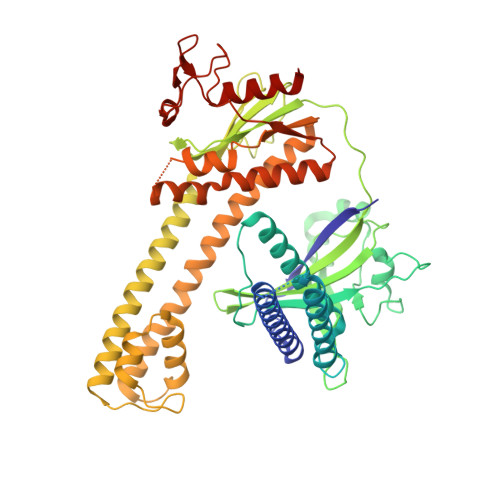
Mechanistic and evolutionary insights into a type V-M CRISPR-Cas effector enzyme.
Omura, S.N., Nakagawa, R., Sudfeld, C., Villegas Warren, R., Wu, W.Y., Hirano, H., Laffeber, C., Kusakizako, T., Kise, Y., Lebbink, J.H.G., Itoh, Y., van der Oost, J., Nureki, O.(2023) Nat Struct Mol Biol 30: 1172-1182
- PubMed: 37460897
- DOI: https://doi.org/10.1038/s41594-023-01042-3
- Primary Citation of Related Structures:
8HHL, 8HHM, 8HIO - PubMed Abstract:
RNA-guided type V CRISPR-Cas12 effectors provide adaptive immunity against mobile genetic elements (MGEs) in bacteria and archaea. Among diverse Cas12 enzymes, the recently identified Cas12m2 (CRISPR-Cas type V-M) is highly compact and has a unique RuvC active site. Although the non-canonical RuvC triad does not permit dsDNA cleavage, Cas12m2 still protects against invading MGEs through transcriptional silencing by strong DNA binding. However, the molecular mechanism of RNA-guided genome inactivation by Cas12m2 remains unknown. Here we report cryo-electron microscopy structures of two states of Cas12m2-CRISPR RNA (crRNA)-target DNA ternary complexes and the Cas12m2-crRNA binary complex, revealing structural dynamics during crRNA-target DNA heteroduplex formation. The structures indicate that the non-target DNA strand is tightly bound to a unique arginine-rich cluster in the recognition (REC) domains and the non-canonical active site in the RuvC domain, ensuring strong DNA-binding affinity of Cas12m2. Furthermore, a structural comparison of Cas12m2 with TnpB, a putative ancestor of Cas12 enzymes, suggests that the interaction of the characteristic coiled-coil REC2 insertion with the protospacer-adjacent motif-distal region of the heteroduplex is crucial for Cas12m2 to engage in adaptive immunity. Collectively, our findings improve mechanistic understanding of diverse type V CRISPR-Cas effectors and provide insights into the evolution of TnpB to Cas12 enzymes.
- Department of Biological Sciences, Graduate School of Science, the University of Tokyo, Tokyo, Japan.
Organizational Affiliation: