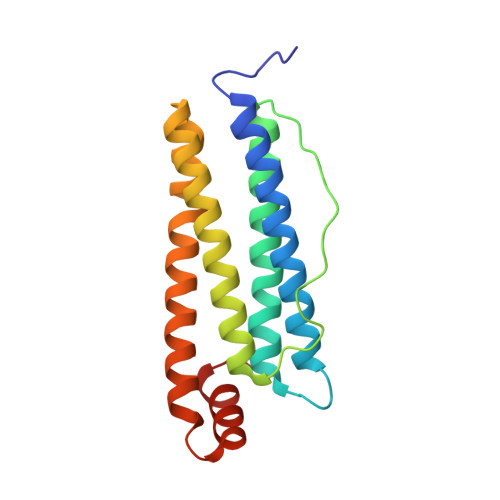
Elucidating Conformational Dynamics and Thermostability of Designed Aromatic Clusters by Using Protein Cages.
Hishikawa, Y., Noya, H., Nagatoishi, S., Yoshidome, T., Maity, B., Tsumoto, K., Abe, S., Ueno, T.(2023) Chemistry 29: e202300488-e202300488
- PubMed: 37070368 Search on PubMed
- DOI: https://doi.org/10.1002/chem.202300488
- Primary Citation Related Structures:
8H8L, 8H8M, 8H8N, 8H8O - PubMed Abstract:
Multiple aromatic residues assemble to form higher ordered structures known as "aromatic clusters" in proteins and play essential roles in biological systems. However, the stabilization mechanism and dynamic behavior of aromatic clusters remain unclear. This study describes designed aromatic interactions confined within a protein cage to reveal how aromatic clusters affect protein stability. The crystal structures and calorimetric measurements indicate that the formation of inter-subunit phenylalanine clusters enhance the interhelix interactions and increase the melting temperature. Theoretical calculations suggest that this is caused by the transformation of the T-shaped geometry into π-π stacking at high temperatures, and the hydration entropic gain. Thus, the isolated nanoenvironment in a protein cage allows reconstruction and detailed analysis of multiple clustering residues for elucidating the mechanisms of various biomolecular interactions in nature which can be applied to design of bionanomaterials.
- School of Life Science and Technology, Tokyo Institute of Technology, Nagatsuta-cho 4259-B55, Midori-ku, 226-8501, Yokohama, Japan.
Organizational Affiliation: