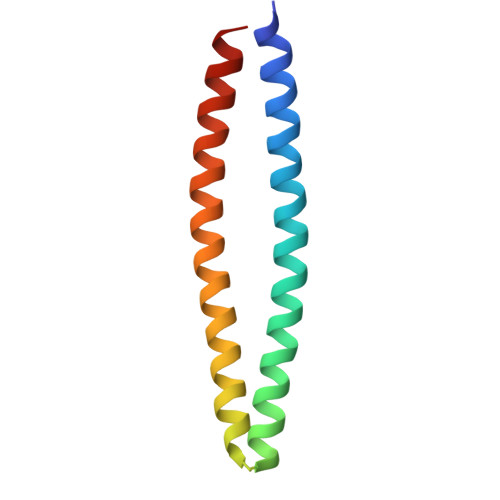Crystal structure and activity of a de novo enzyme, ferric enterobactin esterase Syn-F4.
Kurihara, K., Umezawa, K., Donnelly, A.E., Sperling, B., Liao, G., Hecht, M.H., Arai, R.(2023) Proc Natl Acad Sci U S A 120: e2218281120-e2218281120
- PubMed: 37695900
- DOI: https://doi.org/10.1073/pnas.2218281120
- Primary Citation of Related Structures:
8H7C, 8H7D, 8H7E - PubMed Abstract:
Producing novel enzymes that are catalytically active in vitro and biologically functional in vivo is a key goal of synthetic biology. Previously, we reported Syn-F4, the first de novo protein that meets both criteria. Syn-F4 hydrolyzed the siderophore ferric enterobactin, and expression of Syn-F4 allowed an inviable strain of Escherichia coli (Δ fes ) to grow in iron-limited medium. Here, we describe the crystal structure of Syn-F4. Syn-F4 forms a dimeric 4-helix bundle. Each monomer comprises two long α-helices, and the loops of the Syn-F4 dimer are on the same end of the bundle ( syn topology). Interestingly, there is a penetrated hole in the central region of the Syn-F4 structure. Extensive mutagenesis experiments in a previous study showed that five residues (Glu26, His74, Arg77, Lys78, and Arg85) were essential for enzymatic activity in vivo. All these residues are located around the hole in the central region of the Syn-F4 structure, suggesting a putative active site with a catalytic dyad (Glu26-His74). The complete inactivity of purified proteins with mutations at the five residues supports the putative active site and reaction mechanism. Molecular dynamics and docking simulations of the ferric enterobactin siderophore binding to the Syn-F4 structure demonstrate the dynamic property of the putative active site. The structure and active site of Syn-F4 are completely different from native enterobactin esterase enzymes, thereby demonstrating that proteins designed de novo can provide life-sustaining catalytic activities using structures and mechanisms dramatically different from those that arose in nature.
- Department of Applied Biology, Faculty of Textile Science and Technology, Shinshu University, Ueda, Nagano 386-8567, Japan.
Organizational Affiliation: