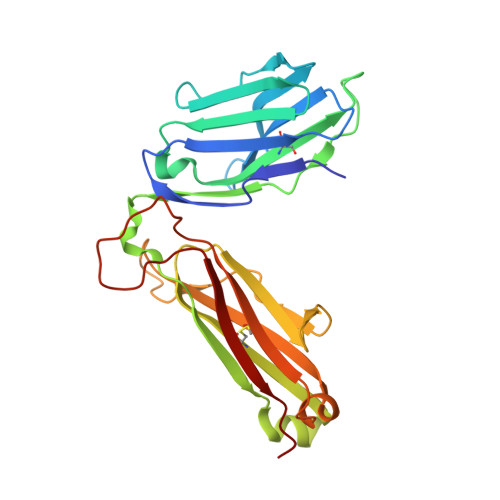
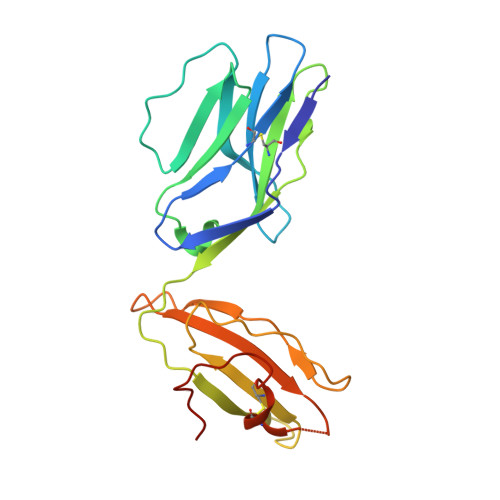
Structural insights into protection against a SARS-CoV-2 spike variant by T cell receptor (TCR) diversity.
Wu, D., Efimov, G.A., Bogolyubova, A.V., Pierce, B.G., Mariuzza, R.A.(2023) J Biological Chem 299: 103035-103035
- PubMed: 36806685 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2023.103035
- Primary Citation Related Structures:
8GOM, 8GON, 8GOP - PubMed Abstract:
T cells play a crucial role in combatting SARS-CoV-2 and forming long-term memory responses to this coronavirus. The emergence of SARS-CoV-2 variants that can evade T cell immunity has raised concerns about vaccine efficacy and the risk of reinfection. Some SARS-CoV-2 T cell epitopes elicit clonally restricted CD8 + T cell responses characterized by T cell receptors (TCRs) that lack structural diversity. Mutations in such epitopes can lead to loss of recognition by most T cells specific for that epitope, facilitating viral escape. Here, we studied an HLA-A2-restricted spike protein epitope (RLQ) that elicits CD8 + T cell responses in COVID-19 convalescent patients characterized by highly diverse TCRs. We previously reported the structure of an RLQ-specific TCR (RLQ3) with greatly reduced recognition of the most common natural variant of the RLQ epitope (T1006I). Opposite to RLQ3, TCR RLQ7 recognizes T1006I with even higher functional avidity than the WT epitope. To explain the ability of RLQ7, but not RLQ3, to tolerate the T1006I mutation, we determined structures of RLQ7 bound to RLQ-HLA-A2 and T1006I-HLA-A2. These complexes show that there are multiple structural solutions to recognizing RLQ and thereby generating a clonally diverse T cell response to this epitope that assures protection against viral escape and T cell clonal loss.
- Laboratory of Structural Immunology, Department of Hepatopancreatobiliary Surgery, The First Affiliated Hospital, Hengyang Medical School, University of South China, Hengyang, Hunan, China; W.M. Keck Laboratory for Structural Biology, University of Maryland Institute for Bioscience and Biotechnology Research, Rockville, Maryland, USA; Department of Cell Biology and Molecular Genetics, University of Maryland, College Park, Maryland, USA.
Organizational Affiliation: