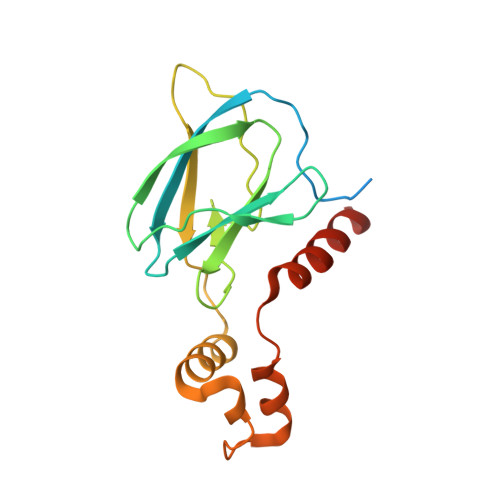
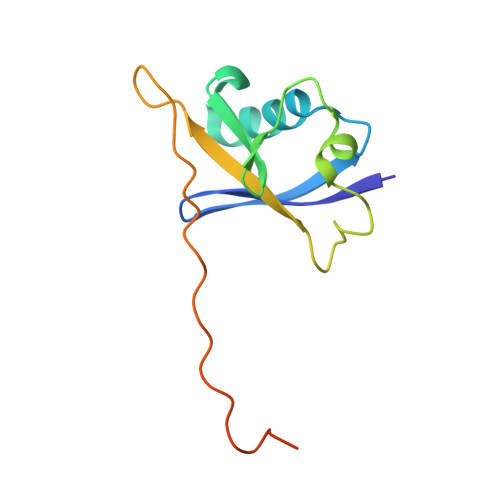
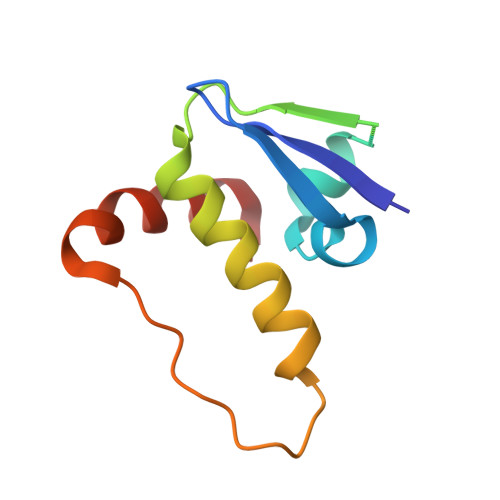
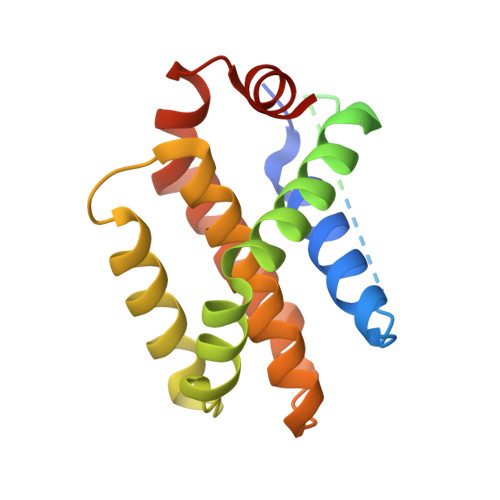
Development and crystal structures of a potent second-generation dual degrader of BCL-2 and BCL-xL.
Nayak, D., Lv, D., Yuan, Y., Zhang, P., Hu, W., Nayak, A., Ruben, E.A., Lv, Z., Sung, P., Hromas, R., Zheng, G., Zhou, D., Olsen, S.K.(2024) Nat Commun 15: 2743-2743
- PubMed: 38548768 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-46922-4
- Primary Citation Related Structures:
8FY0, 8FY1, 8FY2 - PubMed Abstract:
Overexpression of BCL-xL and BCL-2 play key roles in tumorigenesis and cancer drug resistance. Advances in PROTAC technology facilitated recent development of the first BCL-xL/BCL-2 dual degrader, 753b, a VHL-based degrader with improved potency and reduced toxicity compared to previous small molecule inhibitors. Here, we determine crystal structures of VHL/753b/BCL-xL and VHL/753b/BCL-2 ternary complexes. The two ternary complexes exhibit markedly different architectures that are accompanied by distinct networks of interactions at the VHL/753b-linker/target interfaces. The importance of these interfacial contacts is validated via functional analysis and informed subsequent rational and structure-guided design focused on the 753b linker and BCL-2/BCL-xL warhead. This results in the design of a degrader, WH244, with enhanced potency to degrade BCL-xL/BCL-2 in cells. Using biophysical assays followed by in cell activities, we are able to explain the enhanced target degradation of BCL-xL/BCL-2 in cells. Most PROTACs are empirically designed and lack structural studies, making it challenging to understand their modes of action and specificity. Our work presents a streamlined approach that combines rational design and structure-based insights backed with cell-based studies to develop effective PROTAC-based cancer therapeutics.
- Department of Biochemistry & Structural Biology and Greehey Children's Cancer Research Institute, University of Texas Health Science Center at San Antonio, San Antonio, TX, 78229, USA.
Organizational Affiliation: