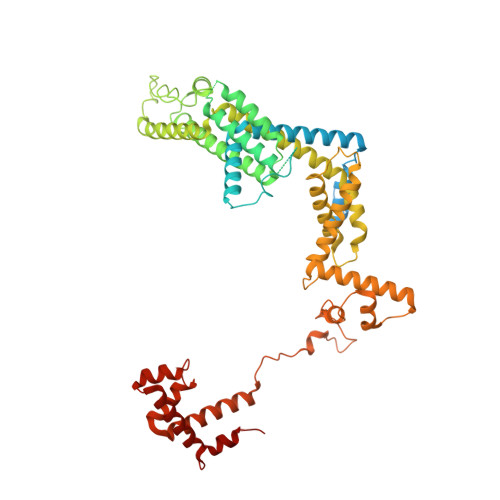
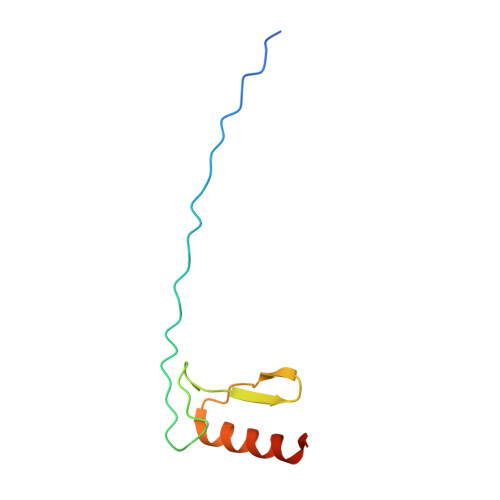
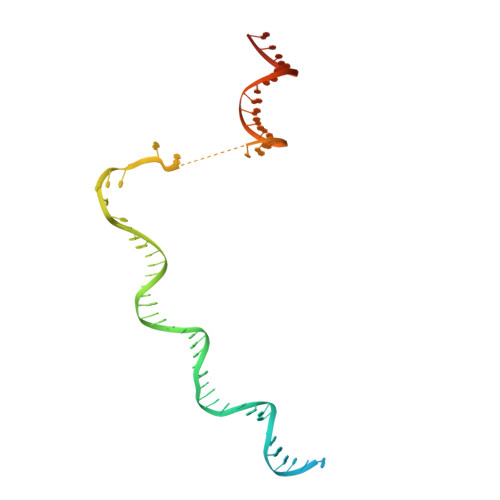
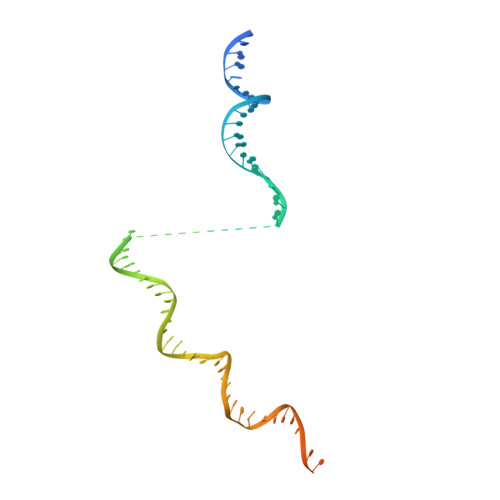
General transcription factor from Escherichia coli with a distinct mechanism of action.
Vasilyev, N., Liu, M.M.J., Epshtein, V., Shamovsky, I., Nudler, E.(2024) Nat Struct Mol Biol 31: 141-149
- PubMed: 38177674 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-023-01154-w
- Primary Citation Related Structures:
8FTD - PubMed Abstract:
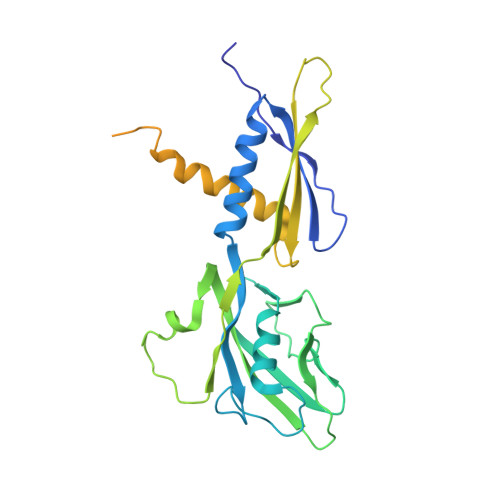
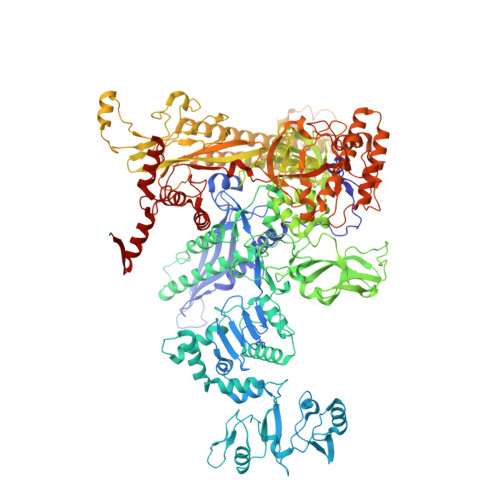
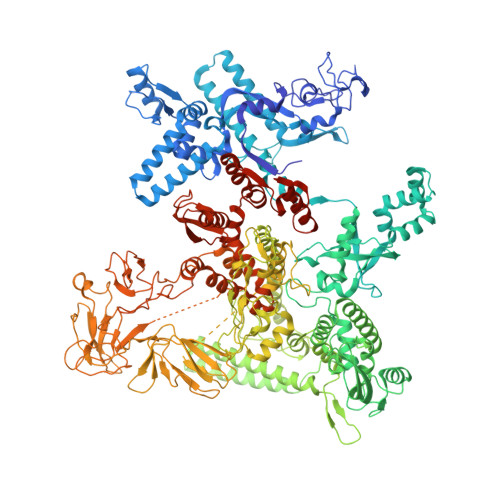
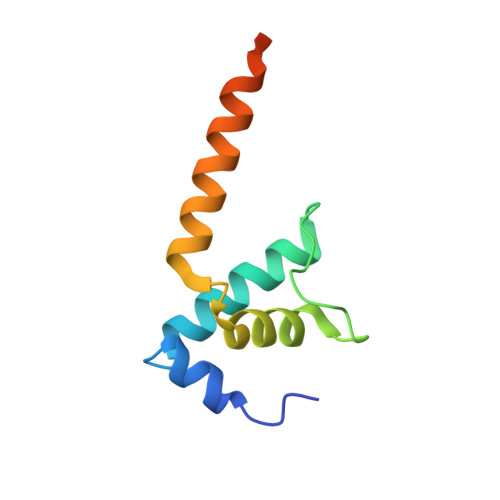
Gene expression in Escherichia coli is controlled by well-established mechanisms that activate or repress transcription. Here, we identify CedA as an unconventional transcription factor specifically associated with the RNA polymerase (RNAP) σ 70 holoenzyme. Structural and biochemical analysis of CedA bound to RNAP reveal that it bridges distant domains of β and σ 70 subunits to stabilize an open-promoter complex. CedA does so without contacting DNA. We further show that cedA is strongly induced in response to amino acid starvation, oxidative stress and aminoglycosides. CedA provides a basal level of tolerance to these clinically relevant antibiotics, as well as to rifampicin and peroxide. Finally, we show that CedA modulates transcription of hundreds of bacterial genes, which explains its pleotropic effect on cell physiology and pathogenesis.
- Department of Biochemistry and Molecular Pharmacology, New York University School of Medicine, New York, NY, USA.
Organizational Affiliation: