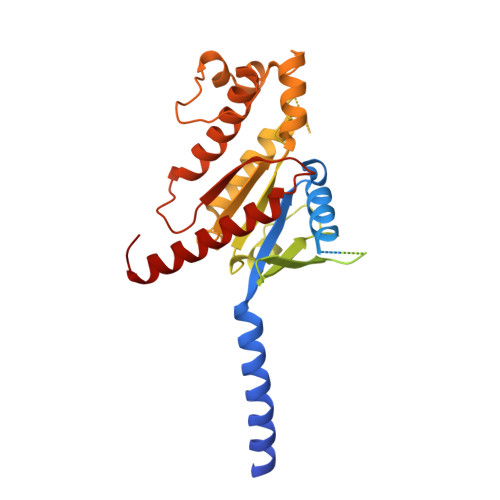
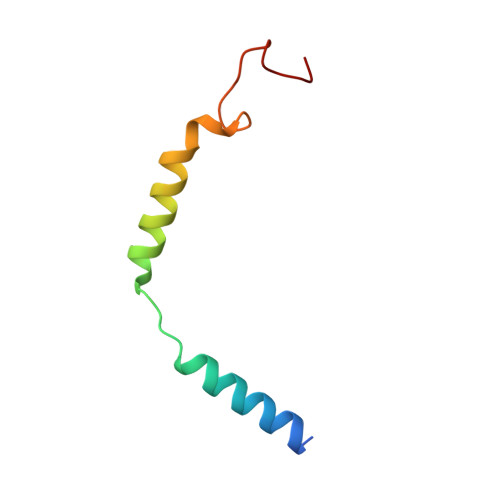
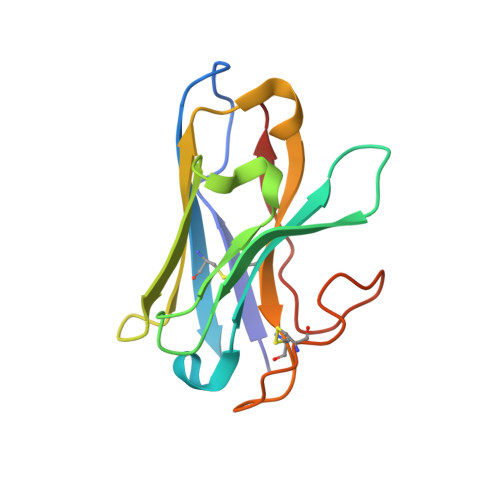
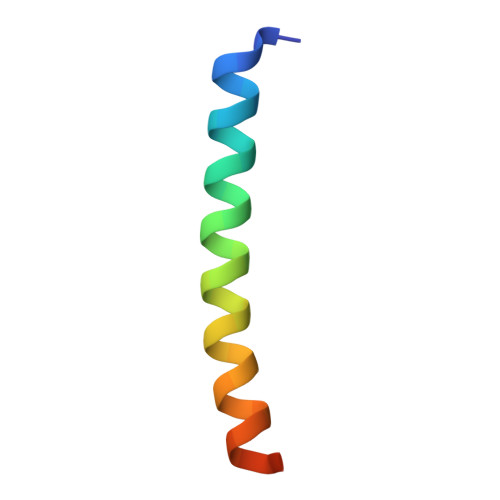
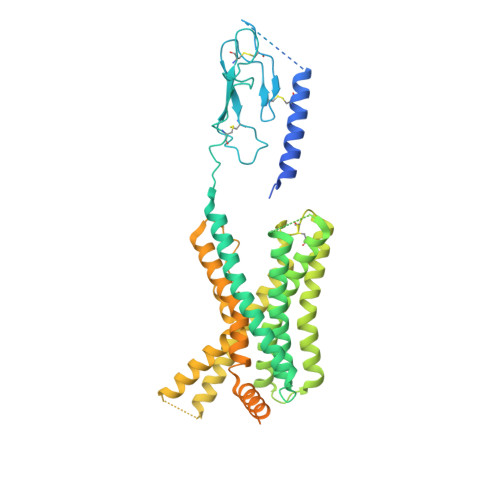
Molecular insights into peptide agonist engagement with the PTH receptor.
Cary, B.P., Gerrard, E.J., Belousoff, M.J., Fletcher, M.M., Jiang, Y., Russell, I.C., Piper, S.J., Wootten, D., Sexton, P.M.(2023) Structure 31: 668-676.e5
- PubMed: 37148874 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2023.04.002
- Primary Citation Related Structures:
8FLQ, 8FLR, 8FLS, 8FLT, 8FLU - PubMed Abstract:
The parathyroid hormone (PTH) 1 receptor (PTH1R) is a G protein-coupled receptor (GPCR) that regulates skeletal development and calcium homeostasis. Here, we describe cryo-EM structures of the PTH1R in complex with fragments of the two hormones, PTH and PTH-related protein, the drug abaloparatide, as well as the engineered tool compounds, long-acting PTH (LA-PTH) and the truncated peptide, M-PTH(1-14). We found that the critical N terminus of each agonist engages the transmembrane bundle in a topologically similar fashion, reflecting similarities in measures of Gαs activation. The full-length peptides induce subtly different extracellular domain (ECD) orientations relative to the transmembrane domain. In the structure bound to M-PTH, the ECD is unresolved, demonstrating that the ECD is highly dynamic when unconstrained by a peptide. High resolutions enabled identification of water molecules near peptide and G protein binding sites. Our results illuminate the action of orthosteric agonists of the PTH1R.
- Drug Discovery Biology Theme, Monash Institute of Pharmaceutical Sciences, Monash University, Parkville 3052, VIC, Australia; ARC Centre for Cryo-Electron Microscopy of Membrane Proteins, Monash Institute of Pharmaceutical Sciences, Monash University, Parkville 3052, VIC, Australia. Electronic address: brian.cary@monash.edu.
Organizational Affiliation: