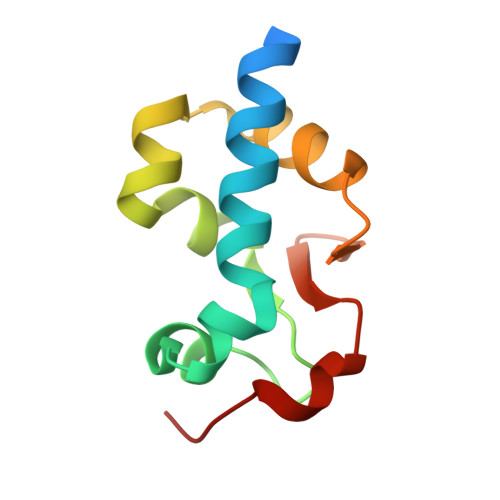
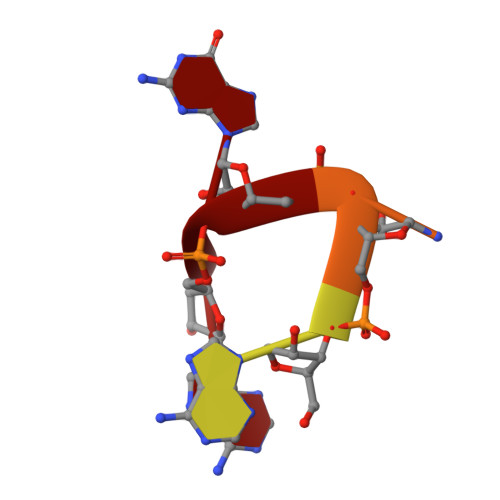
Enhanced binding of guanylated poly(A) RNA by the LaM domain of LARP1.
Kozlov, G., Jiang, J., Rutherford, T., Noronha, A.M., Wilds, C.J., Gehring, K.(2024) RNA Biol 21: 7-16
- PubMed: 39016322 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1080/15476286.2024.2379121
- Primary Citation Related Structures:
7SOW, 8EY6, 8EY7, 8EY8, 8G90, 8G91 - PubMed Abstract:
La-related proteins (LARPs) are a family of RNA-binding proteins that share a conserved La motif (LaM) domain. LARP1 plays a role in regulating ribosomal protein synthesis and stabilizing mRNAs and has a unique structure without an RNA binding RRM domain adjoining the LaM domain. In this study, we investigated the physical basis for LARP1 specificity for poly(A) sequences and observed an unexpected bias for sequences with single guanines. Multiple guanine substitutions did not increase the affinity, demonstrating preferential recognition of singly guanylated sequences. We also observed that the cyclic di-nucleotides in the cCAS/STING pathway, cyclic-di-GMP and 3',3'-cGAMP, bound with sub-micromolar affinity. Isothermal titration measurements were complemented by high-resolution crystal structures of the LARP1 LaM with six different RNA ligands, including two stereoisomers of a phosphorothioate linkage. The selectivity for singly substituted poly(A) sequences suggests LARP1 may play a role in the stabilizing effect of poly(A) tail guanylation. [Figure: see text].
- Department of Biochemistry, McGill University, Montréal, Quebec, Canada.
Organizational Affiliation: