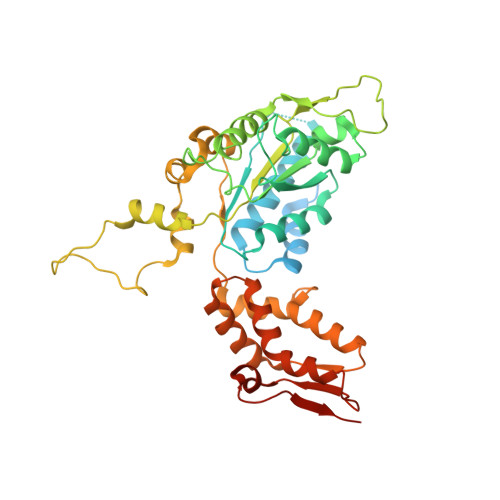
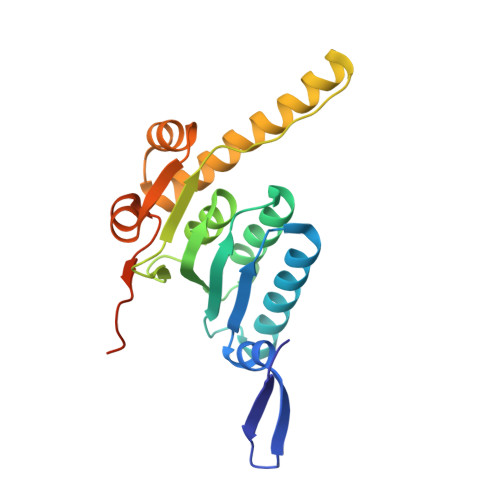
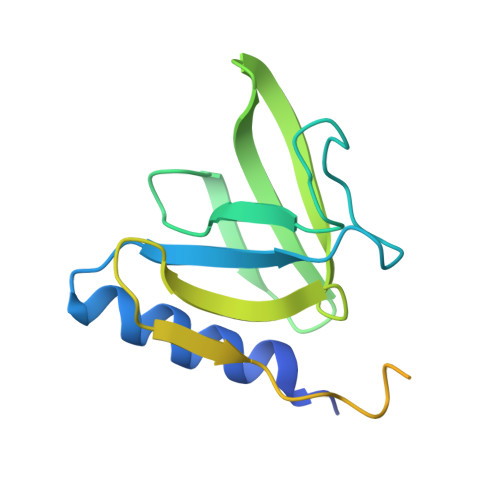
The SspB adaptor drives structural changes in the AAA+ ClpXP protease during ssrA-tagged substrate delivery.
Ghanbarpour, A., Fei, X., Baker, T.A., Davis, J.H., Sauer, R.T.(2023) Proc Natl Acad Sci U S A 120: e2219044120-e2219044120
- PubMed: 36730206 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2219044120
- Primary Citation Related Structures:
8ET3 - PubMed Abstract:
Energy-dependent protein degradation by the AAA+ ClpXP protease helps maintain protein homeostasis in bacteria and eukaryotic organelles of bacterial origin. In Escherichia coli and many other proteobacteria, the SspB adaptor assists ClpXP in degrading ssrA-tagged polypeptides produced as a consequence of tmRNA-mediated ribosome rescue. By tethering these incomplete ssrA-tagged proteins to ClpXP, SspB facilitates their efficient degradation at low substrate concentrations. How this process occurs structurally is unknown. Here, we present a cryo-EM structure of the SspB adaptor bound to a GFP-ssrA substrate and to ClpXP. This structure provides evidence for simultaneous contacts of SspB and ClpX with the ssrA tag within the tethering complex, allowing direct substrate handoff concomitant with the initiation of substrate translocation. Furthermore, our structure reveals that binding of the substrate·adaptor complex induces unexpected conformational changes within the spiral structure of the AAA+ ClpX hexamer and its interaction with the ClpP tetradecamer.
- Department of Biology, Massachusetts Institute of Technology, Cambridge, MA 02139.
Organizational Affiliation: