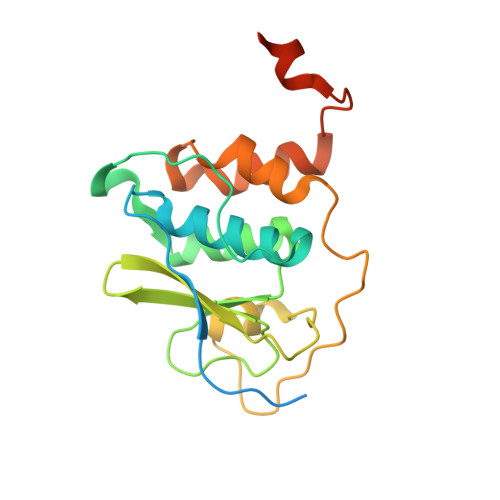
Demonstrating the importance of porcine reproductive and respiratory syndrome virus papain-like protease 2 deubiquitinating activity in viral replication by structure-guided mutagenesis.
Bailey-Elkin, B.A., Knaap, R.C.M., De Silva, A., Boekhoud, I.M., Mous, S., van Vught, N., Khajehpour, M., van den Born, E., Kikkert, M., Mark, B.L.(2023) PLoS Pathog 19: e1011872-e1011872
- PubMed: 38096325 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1011872
- Primary Citation Related Structures:
8EHN, 8EHO - PubMed Abstract:
Deubiquitination of cellular substrates by viral proteases is a mechanism used to interfere with host cellular signaling processes, shared between members of the coronavirus- and arterivirus families. In the case of Arteriviruses, deubiquitinating and polyprotein processing activities are accomplished by the virus-encoded papain-like protease 2 (PLP2). Several studies have implicated the deubiquitinating activity of the porcine reproductive and respiratory syndrome virus (PRRSV) PLP2 in the downregulation of cellular interferon production, however to date, the only arterivirus PLP2 structure described is that of equine arteritis virus (EAV), a distantly related virus. Here we describe the first crystal structure of the PRRSV PLP2 domain both in the presence and absence of its ubiquitin substrate, which reveals unique structural differences in this viral domain compared to PLP2 from EAV. To probe the role of PRRSV PLP2 deubiquitinating activity in host immune evasion, we selectively removed this activity from the domain by mutagenesis and found that the viral domain could no longer downregulate cellular interferon production. Interestingly, unlike EAV, and also unlike the situation for MERS-CoV, we found that recombinant PRRSV carrying PLP2 DUB-specific mutations faces significant selective pressure to revert to wild-type virus in MARC-145 cells, suggesting that the PLP2 DUB activity, which in PRRSV is present as three different versions of viral protein nsp2 expressed during infection, is critically important for PRRSV replication.
- Department of Microbiology, University of Manitoba, Winnipeg, Manitoba, Canada.
Organizational Affiliation: