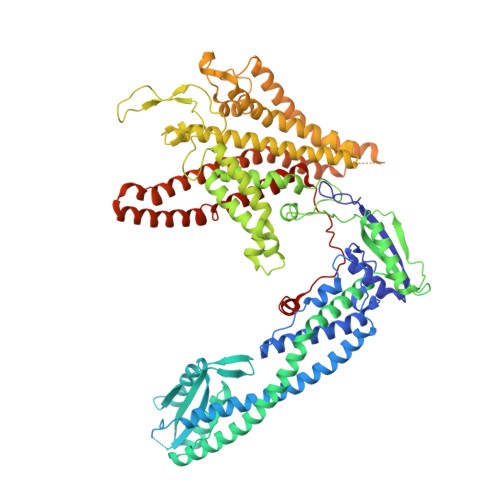
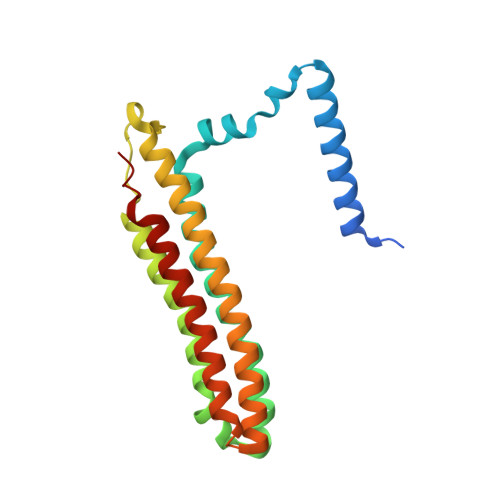
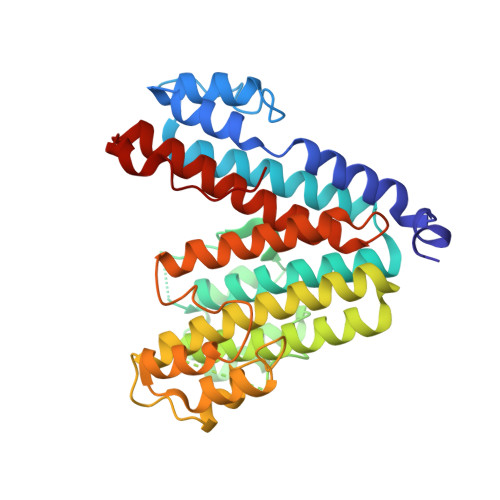
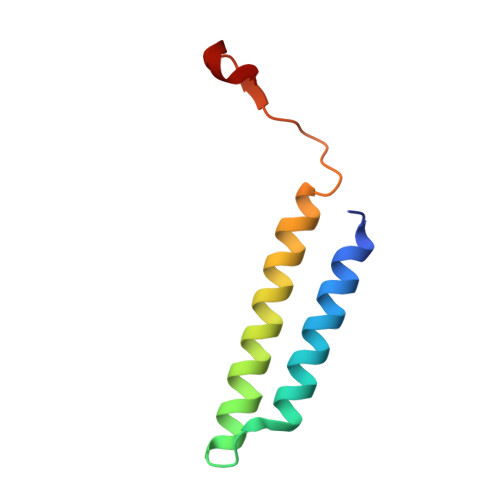
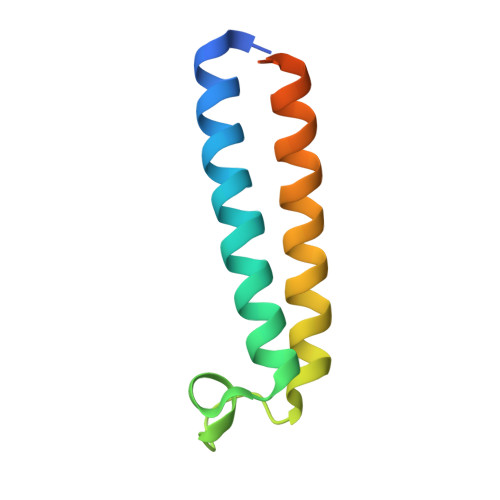
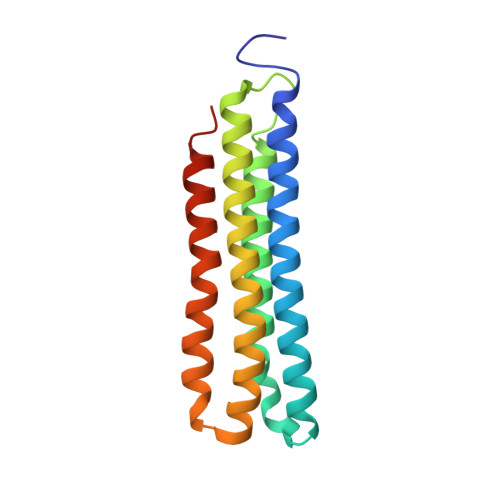
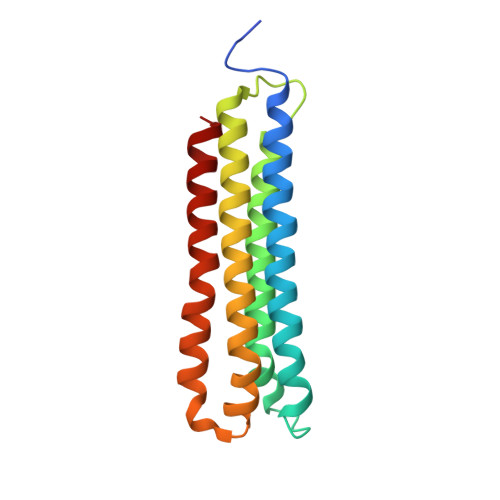
Structural basis of V-ATPase V O region assembly by Vma12p, 21p, and 22p.
Wang, H., Bueler, S.A., Rubinstein, J.L.(2023) Proc Natl Acad Sci U S A 120: e2217181120-e2217181120
- PubMed: 36724250 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2217181120
- Primary Citation Related Structures:
8EAS, 8EAT, 8EAU, 8EAV - PubMed Abstract:
Vacuolar-type adenosine triphosphatases (V-ATPases) are rotary proton pumps that acidify specific intracellular compartments in almost all eukaryotic cells. These multi-subunit enzymes consist of a soluble catalytic V 1 region and a membrane-embedded proton-translocating V O region. V O is assembled in the endoplasmic reticulum (ER) membrane, and V 1 is assembled in the cytosol. However, V 1 binds V O only after V O is transported to the Golgi membrane, thereby preventing acidification of the ER. We isolated V O complexes and subcomplexes from Saccharomyces cerevisiae bound to V-ATPase assembly factors Vma12p, Vma21p, and Vma22p. Electron cryomicroscopy shows how the Vma12-22p complex recruits subunits a, e, and f to the rotor ring of V O while blocking premature binding of V 1 . Vma21p, which contains an ER-retrieval motif, binds the V O :Vma12-22p complex, "mature" V O , and a complex that appears to contain a ring of loosely packed rotor subunits and the proteins YAR027W and YAR028W. The structures suggest that Vma21p binds assembly intermediates that contain a rotor ring and that activation of proton pumping following assembly of V 1 with V O removes Vma21p, allowing V-ATPase to remain in the Golgi. Together, these structures show how Vma12-22p and Vma21p function in V-ATPase assembly and quality control, ensuring the enzyme acidifies only its intended cellular targets.
- Molecular Medicine Program, The Hospital for Sick Children, Toronto M5G 0A4, Canada.
Organizational Affiliation: