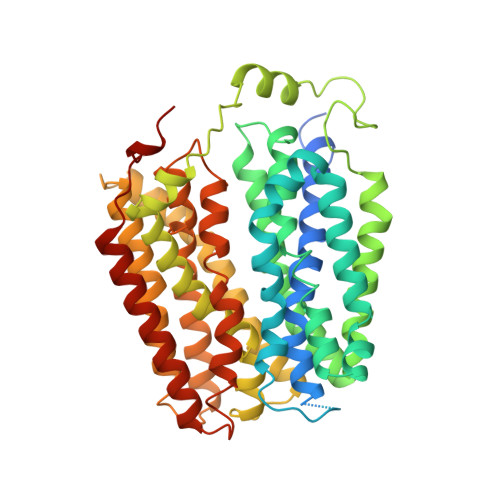
The molecular mechanism of sialic acid transport mediated by Sialin.
Hu, W., Chi, C., Song, K., Zheng, H.(2023) Sci Adv 9: eade8346-eade8346
- PubMed: 36662855 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.ade8346
- Primary Citation Related Structures:
8DWI - PubMed Abstract:
Malfunction of the sialic acid transporter caused by various genetic mutations in the SLC17A5 gene encoding Sialin leads to a spectrum of neurodegenerative conditions called free sialic acid storage disorders. Unfortunately, how Sialin transports sialic acid/proton (H + ) and how pathogenic mutations impair its function are poorly defined. Here, we present the structure of human Sialin in an inward-facing partially open conformation determined by cryo-electron microscopy, representing the first high-resolution structure of any human SLC17 member. Our analysis reveals two unique features in Sialin: (i) The H + coupling/sensing requires two highly conserved Glu residues (E171 and E175) instead of one (E175) as implied in previous studies; and (ii) the normal function of Sialin requires the stabilization of a cytosolic helix, which has not been noticed in the literature. By mapping known pathogenic mutations, we provide mechanistic explanations for corresponding functional defects. We propose a structure-based mechanism for sialic acid transport mediated by Sialin.
- Department of Biochemistry and Molecular Genetics, University of Colorado Anschutz Medical Campus, School of Medicine, Aurora, CO, USA.
Organizational Affiliation: