Native serotonin transporter in complex with 15B8 Fab in the presence of methamphetamine
Yang, D., Gouaux, E.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Transporter | 539 | Sus scrofa | Mutation(s): 0 Membrane Entity: Yes | 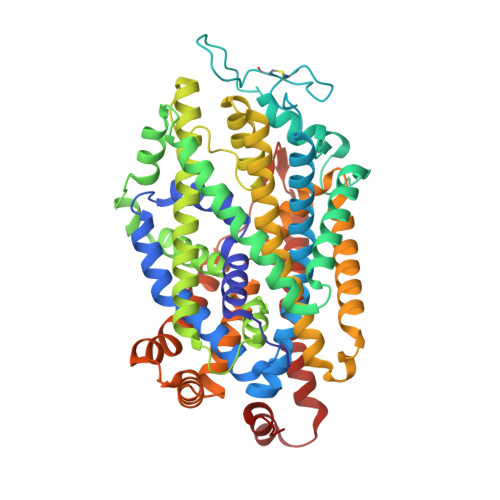 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | F1RN70 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | |||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 15B8 Fab heavy chain variable domain | 118 | Mus musculus | Mutation(s): 0 | 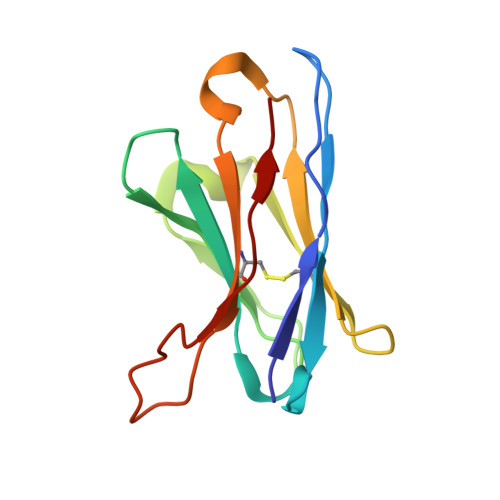 | |
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 15B8 Fab light chain variable domain | 110 | Mus musculus | Mutation(s): 0 | 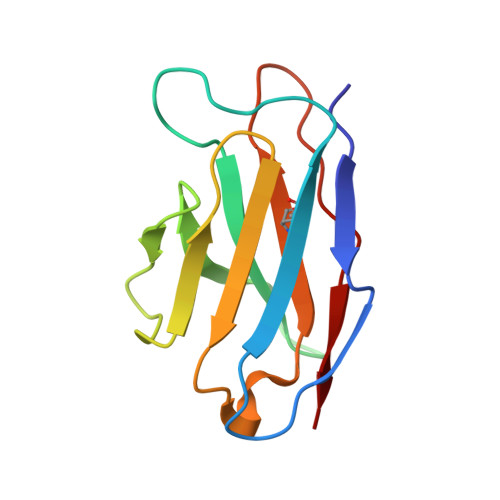 | |
| Ligands 5 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| LMT (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | K [auth A] | DODECYL-BETA-D-MALTOSIDE C24 H46 O11 NLEBIOOXCVAHBD-QKMCSOCLSA-N |  | ||
| Y01 Download:Ideal Coordinates CCD File | I [auth A], J [auth A] | CHOLESTEROL HEMISUCCINATE C31 H50 O4 WLNARFZDISHUGS-MIXBDBMTSA-N |  | ||
| B40 (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | H [auth A] | (2S)-N-methyl-1-phenylpropan-2-amine C10 H15 N MYWUZJCMWCOHBA-VIFPVBQESA-N |  | ||
| CL Download:Ideal Coordinates CCD File | E [auth A] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| NA Download:Ideal Coordinates CCD File | F [auth A], G [auth A] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Howard Hughes Medical Institute (HHMI) | United States | -- |
| National Institutes of Health/National Center for Research Resources (NIH/NCRR) | United States | -- |