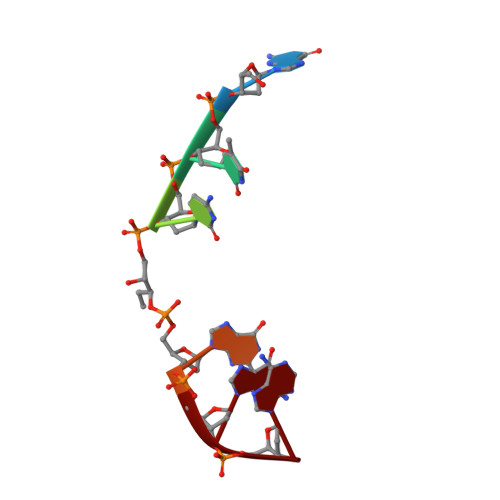
The SOS response-associated peptidase (SRAP) domain of YedK catalyzes ring opening of abasic sites and reversal of its DNA-protein cross-link.
Paulin, K.A., Cortez, D., Eichman, B.F.(2022) J Biological Chem 298: 102307-102307
- PubMed: 35934051 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2022.102307
- Primary Citation Related Structures:
8D2M - PubMed Abstract:
Apurinic/apyrimidinic (AP, or abasic) sites in DNA are one of the most common forms of DNA damage. AP sites are reactive and form cross-links to both proteins and DNA, are prone to strand breakage, and inhibit DNA replication and transcription. The replication-associated AP site repair protein HMCES protects cells from strand breaks, inhibits mutagenic translesion synthesis, and participates in repair of interstrand DNA cross-links derived from AP sites by forming a stable thiazolidine DNA-protein cross-link (DPC) to AP sites in single-stranded DNA (ssDNA). Despite the importance of HMCES to genome maintenance and the evolutionary conservation of its catalytic SRAP (SOS Response Associated Peptidase) domain, the enzymatic mechanisms of DPC formation and resolution are unknown. Using the bacterial homolog YedK, we show that the SRAP domain catalyzes conversion of the AP site to its reactive, ring-opened aldehyde form, and we provide structural evidence for the Schiff base intermediate that forms prior to the more stable thiazolidine. We also report two new activities, whereby SRAP reacts with polyunsaturated aldehydes at DNA 3'-ends generated by bifunctional DNA glycosylases and catalyzes direct reversal of the DPC to regenerate the AP site, the latter of which we observe in both YedK and HMCES-SRAP proteins. Taken together, this work provides insights into possible mechanisms by which HMCES DPCs are resolved in cells.
- Department of Biological Sciences, Vanderbilt University, Nashville, Tennessee, USA.
Organizational Affiliation: