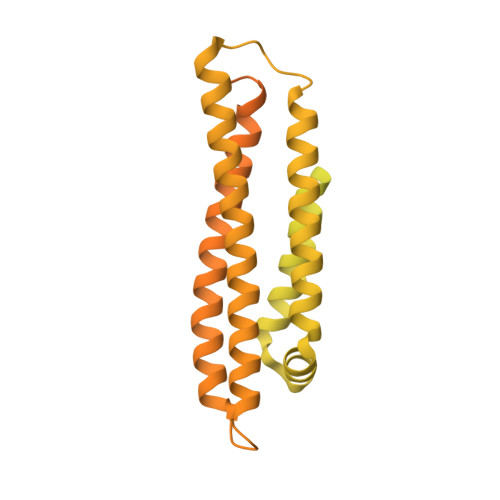
Domino-like effect of C112R mutation on ApoE4 aggregation and its reduction by Alzheimer's Disease drug candidate.
Nemergut, M., Marques, S.M., Uhrik, L., Vanova, T., Nezvedova, M., Gadara, D.C., Jha, D., Tulis, J., Novakova, V., Planas-Iglesias, J., Kunka, A., Legrand, A., Hribkova, H., Pospisilova, V., Sedmik, J., Raska, J., Prokop, Z., Damborsky, J., Bohaciakova, D., Spacil, Z., Hernychova, L., Bednar, D., Marek, M.(2023) Mol Neurodegener 18: 38-38
- PubMed: 37280636 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/s13024-023-00620-9
- Primary Citation Related Structures:
8AX8, 8AX9, 8CDY, 8CE0 - PubMed Abstract:
Apolipoprotein E (ApoE) ε4 genotype is the most prevalent risk factor for late-onset Alzheimer's Disease (AD). Although ApoE4 differs from its non-pathological ApoE3 isoform only by the C112R mutation, the molecular mechanism of its proteinopathy is unknown. Here, we reveal the molecular mechanism of ApoE4 aggregation using a combination of experimental and computational techniques, including X-ray crystallography, site-directed mutagenesis, hydrogen-deuterium mass spectrometry (HDX-MS), static light scattering and molecular dynamics simulations. Treatment of ApoE ε3/ε3 and ε4/ε4 cerebral organoids with tramiprosate was used to compare the effect of tramiprosate on ApoE4 aggregation at the cellular level. We found that C112R substitution in ApoE4 induces long-distance (> 15 Å) conformational changes leading to the formation of a V-shaped dimeric unit that is geometrically different and more aggregation-prone than the ApoE3 structure. AD drug candidate tramiprosate and its metabolite 3-sulfopropanoic acid induce ApoE3-like conformational behavior in ApoE4 and reduce its aggregation propensity. Analysis of ApoE ε4/ε4 cerebral organoids treated with tramiprosate revealed its effect on cholesteryl esters, the storage products of excess cholesterol. Our results connect the ApoE4 structure with its aggregation propensity, providing a new druggable target for neurodegeneration and ageing.
- Loschmidt Laboratories, Department of Experimental Biology, Faculty of Science, Masaryk University, Kamenice 5, Brno, 625 00, Czech Republic.
Organizational Affiliation: