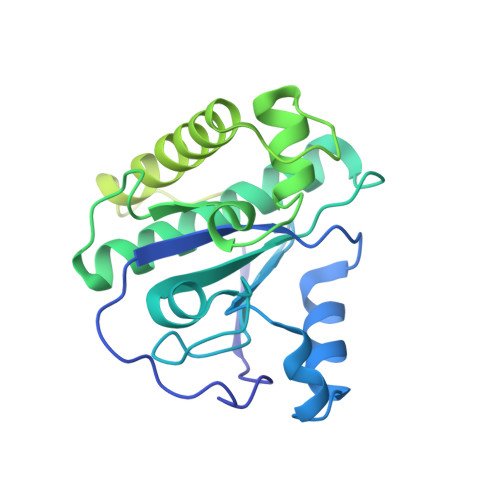
Structural and functional features of a broad-spectrum prophage-encoded enzybiotic from Enterococcus faecium.
Premetis, G.E., Stathi, A., Papageorgiou, A.C., Labrou, N.E.(2023) Sci Rep 13: 7450-7450
- PubMed: 37156923 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-023-34309-2
- Primary Citation Related Structures:
8C4D - PubMed Abstract:
Multidrug-resistant (MDR) bacteria have become a growing threat to public health. The gram-positive Enterococcus faecium is classified by WHO as a high-priority pathogen among the global priority list of antibiotic-resistant bacteria. Peptidoglycan-degrading enzymes (PDEs), also known as enzybiotics, are useful bactericidal agents in the fight against resistant bacteria. In this work, a genome-based screening approach of the genome of E. faecium allowed the identification of a putative PDE gene with predictive amidase activity (EfAmi1; EC 3.5.1.28) in a prophage-integrated sequence. EfAmi1 is composed by two domains: a N-terminal Zn 2+ -dependent N-acetylmuramoyl-L-alanine amidase-2 (NALAA-2) domain and a C-terminal domain with unknown structure and function. The full-length gene of EfAmi1 was cloned and expressed as a 6xHis-tagged protein in E. coli. EfAmi1 was produced as a soluble protein, purified, and its lytic and antimicrobial activities were investigated using turbidity reduction and Kirby-Bauer disk-diffusion assays against clinically isolated bacterial pathogens. The crystal structure of the N-terminal amidase-2 domain was determined using X-ray crystallography at 1.97 Å resolution. It adopts a globular fold with several α-helices surrounding a central five-stranded β-sheet. Sequence comparison revealed a cluster of conserved amino acids that defines a putative binding site for a buried zinc ion. The results of the present study suggest that EfAmi1 displays high lytic and antimicrobial activity and may represent a promising new antimicrobial in the post-antibiotic era.
- Laboratory of Enzyme Technology, Department of Biotechnology, School of Applied Biology and Biotechnology, Agricultural University of Athens, 75 Iera Odos Street, 11855, Athens, Greece.
Organizational Affiliation: