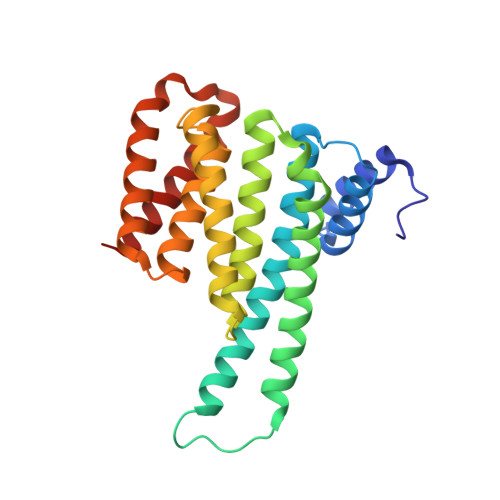
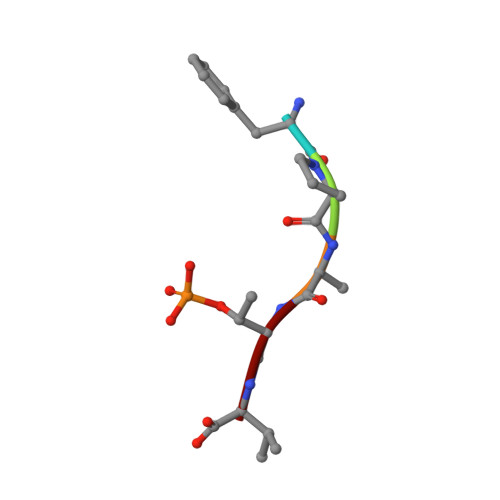
Stabilization of the Estrogen receptor alpha - 14-3-3 interaction as a potential intervention strategy for endocrine resistance in breast cancer
Visser, E.J., Donaldson Collier, M., Siefert, J., Konstantinidou, M., Paul, S., Cossar, P., Miley, G., Meijer, O., Arkin, M.R., Ottmann, C., Zwart, W., Brunsveld, L.To be published.