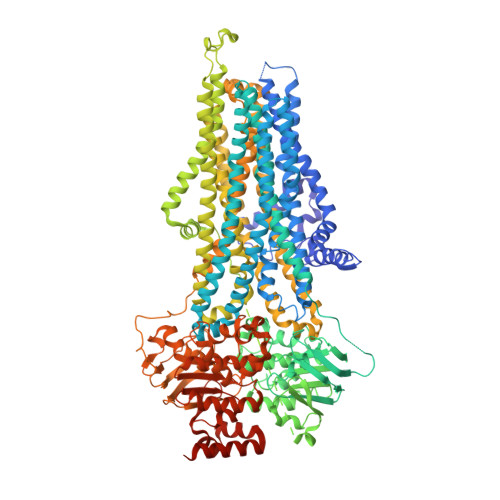
Structural and mechanistic basis of substrate transport by the multidrug transporter MRP4.
Bloch, M., Raj, I., Pape, T., Taylor, N.M.I.(2023) Structure 31: 1407-1418.e6
- PubMed: 37683641 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2023.08.014
- Primary Citation Related Structures:
8BJF, 8BWO, 8BWP, 8BWQ, 8BWR - PubMed Abstract:
Multidrug resistance-associated protein 4 (MRP4) is an ATP-binding cassette (ABC) transporter expressed at multiple tissue barriers where it actively extrudes a wide variety of drug compounds. Overexpression of MRP4 provides resistance to clinically used antineoplastic agents, making it a highly attractive therapeutic target for countering multidrug resistance. Here, we report cryo-EM structures of multiple physiologically relevant states of lipid bilayer-embedded human MRP4, including complexes between MRP4 and two widely used chemotherapeutic agents and a complex between MRP4 and its native substrate. The structures display clear similarities and distinct differences in the coordination of these chemically diverse substrates and, in combination with functional and mutational analysis, reveal molecular details of the transport mechanism. Our study provides key insights into the unusually broad substrate specificity of MRP4 and constitutes an important contribution toward a general understanding of multidrug transporters.
- Structural Biology of Molecular Machines Group, Protein Structure & Function Program, Novo Nordisk Foundation Center for Protein Research, Faculty of Health and Medical Sciences, University of Copenhagen, Blegdamsvej 3B, 2200 Copenhagen, Denmark.
Organizational Affiliation: