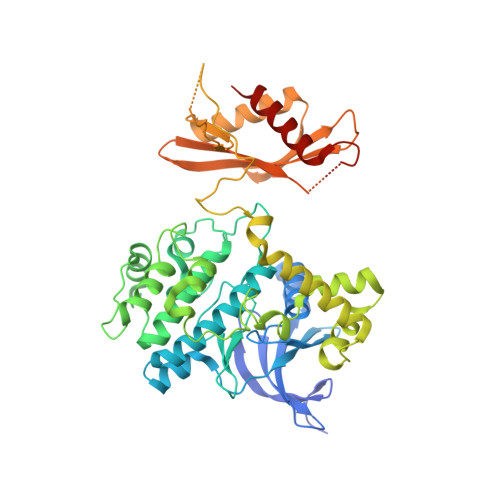
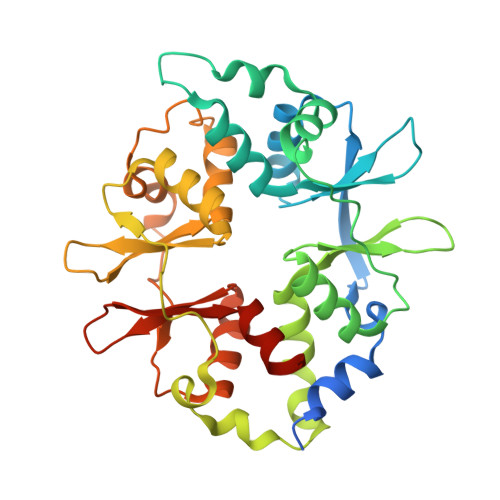
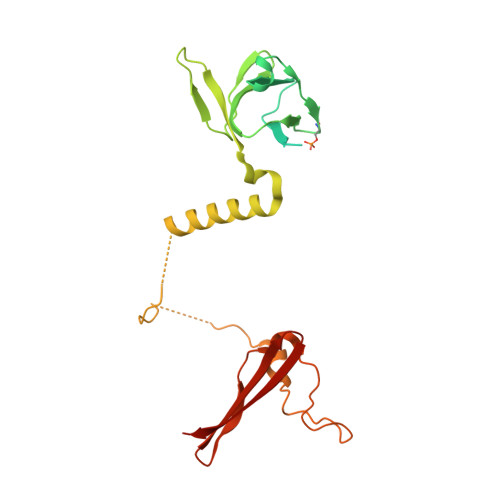Direct beta 1/ beta 2 AMPK activation reduces liver steatosis but not fibrosis in a mouse model of non-alcoholic steatohepatitis
Mather, K.M., Boland, M.L., Rivers, E.L., Srivastava, A., Schimpl, M., Hemsley, P., Robinson, J., Wan, P., Hansen, J., Read, J.A., Trevaskis, J.L., Smith, D.M.(2024) bioRxiv