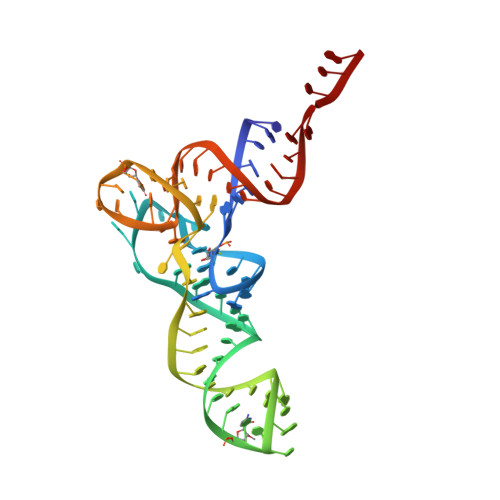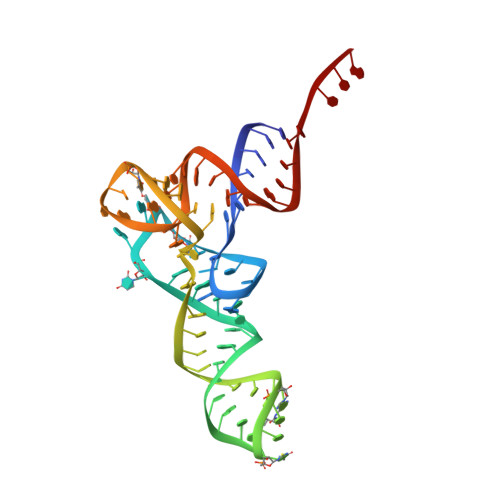Modulation of translational decoding by m 6 A modification of mRNA.
Jain, S., Koziej, L., Poulis, P., Kaczmarczyk, I., Gaik, M., Rawski, M., Ranjan, N., Glatt, S., Rodnina, M.V.(2023) Nat Commun 14: 4784-4784
- PubMed: 37553384 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-40422-7
- Primary Citation Related Structures:
8BF7, 8BGE, 8BGH, 8BH4, 8BHJ, 8BHL, 8BHN, 8BHP, 8BIL, 8BIM - PubMed Abstract:
N 6 -methyladenosine (m 6 A) is an abundant, dynamic mRNA modification that regulates key steps of cellular mRNA metabolism. m 6 A in the mRNA coding regions inhibits translation elongation. Here, we show how m 6 A modulates decoding in the bacterial translation system using a combination of rapid kinetics, smFRET and single-particle cryo-EM. We show that, while the modification does not impair the initial binding of aminoacyl-tRNA to the ribosome, in the presence of m 6 A fewer ribosomes complete the decoding process due to the lower stability of the complexes and enhanced tRNA drop-off. The mRNA codon adopts a π-stacked codon conformation that is remodeled upon aminoacyl-tRNA binding. m 6 A does not exclude canonical codon-anticodon geometry, but favors alternative more dynamic conformations that are rejected by the ribosome. These results highlight how modifications outside the Watson-Crick edge can still interfere with codon-anticodon base pairing and complex recognition by the ribosome, thereby modulating the translational efficiency of modified mRNAs.
- Max Planck Institute for Multidisciplinary Sciences, Göttingen, 37077, Germany.
Organizational Affiliation: