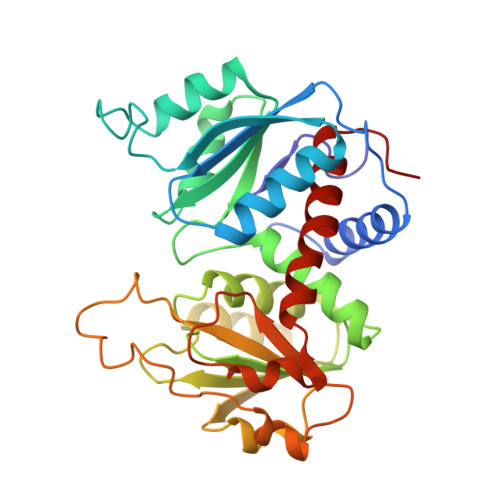
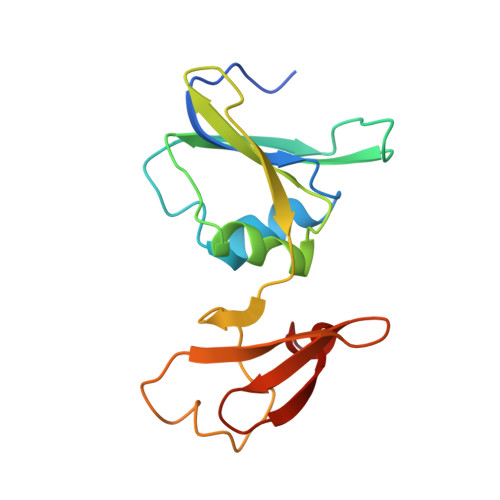
Crystal structures of aspartate carbamoyltransferase ligated with phosphonoacetamide, malonate, and CTP or ATP at 2.8-A resolution and neutral pH.
Gouaux, J.E., Stevens, R.C., Lipscomb, W.N.(1990) Biochemistry 29: 7702-7715
- PubMed: 2271529 Search on PubMed
- DOI: https://doi.org/10.1021/bi00485a020
- Primary Citation Related Structures:
7AT1, 8AT1 - PubMed Abstract:
The R-state structures of the ATP and CTP complexes of aspartate carbamoyltransferase ligated with phosphonoacetamide and malonate have been determined at 2.8-A resolution and neutral pH. These structures were solved by the method of molecular replacement and were refined to crystallographic residuals between 0.167 and 0.182. The triphosphate, the ribose, and the purine and pyrimidine moieties of ATP and CTP interact with similar regions of the allosteric domain of the regulatory dimer. ATP and CTP relatively increase and decrease the size of the allosteric site in the vicinity of the base, respectively. For both CTP and ATP at pH 7, the gamma-phosphates are bound to His20 and are also near Lys94, while the alpha-phosphates interact exclusively with Lys94. The 2'-hydroxyls of both CTP and ATP are near the amino group of Lys60. The pyrimidine ring of CTP makes specific hydrogen bonds at the allosteric site: the NH2 group donates hydrogen bonds to the main-chain carbonyls of Ile12 and Tyr89 and the pyrimidine ring carbonyl oxygen accepts a hydrogen bond from the amino group of Lys60; the nitrogen at position 3 in the pyrimidine ring is hydrogen bonded to a main-chain NH group of Ile12. The purine ring of ATP also makes numerous interactions with residues at the allosteric site: the purine NH2 (analogous to the amino group of CTP) donates a hydrogen bond to the main-chain carbonyl oxygen of Ile12, the N3 nitrogen interacts with the amino group of Lys60, and the N1 nitrogen hydrogen bonds to the NH group of Ile12. The binding of CTP and ATP to the allosteric site in the presence of phosphonoacetamide and malonate does not dramatically alter the structure of the allosteric binding site or of the allosteric domain. Nonetheless, in the CTP-ligated structure, the average separation between the catalytic trimers decreases by approximately 0.5 A, indicating a small shift of the quaternary structure toward the T state. In the CTP- and ATP-ligated R-state structures, the binding and occupancy of phosphonoacetamide and malonate are similar and the structures of the active sites are similar at the current resolution of 2.8 A.
- Gibbs Chemical Laboratory, Harvard University, Cambridge, Massachusetts 02138.
Organizational Affiliation: