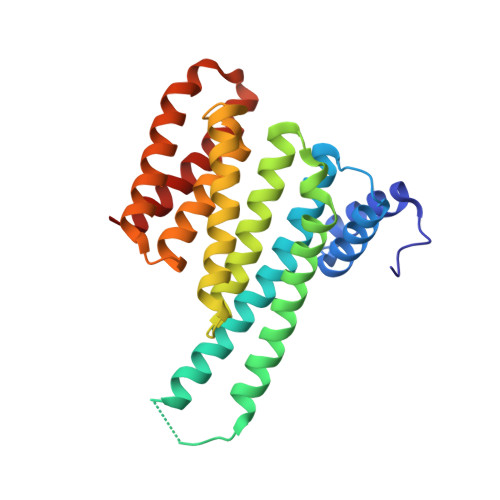
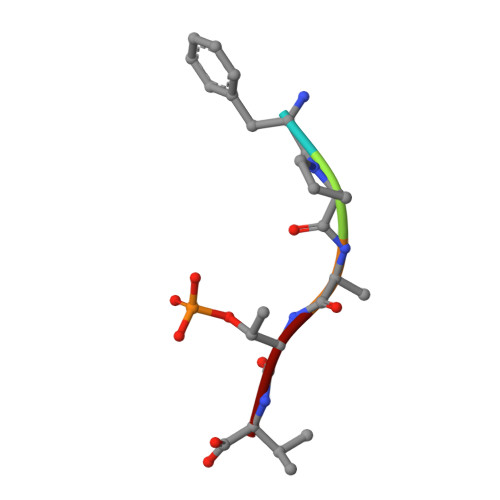
Structure-Based Optimization of Covalent, Small-Molecule Stabilizers of the 14-3-3 sigma /ER alpha Protein-Protein Interaction from Nonselective Fragments.
Konstantinidou, M., Visser, E.J., Vandenboorn, E., Chen, S., Jaishankar, P., Overmans, M., Dutta, S., Neitz, R.J., Renslo, A.R., Ottmann, C., Brunsveld, L., Arkin, M.R.(2023) J Am Chem Soc 145: 20328-20343
- PubMed: 37676236 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.3c05161
- Primary Citation Related Structures:
8AI0, 8ALR, 8ALT, 8ALV, 8ALW, 8AM7, 8ANF, 8AOY, 8APS, 8AQ1, 8AQC, 8AQE, 8AQZ, 8AR4, 8AR5, 8ARG, 8ARO, 8ARQ, 8ARR, 8ARW, 8ARX, 8ARY, 8ARZ, 8AS1, 8AT9, 8ATP, 8ATR, 8ATS, 8AU2, 8AUS, 8AUY, 8AV3, 8AV4, 8AV7, 8AV8, 8AWG, 8AXE, 8AXU, 8AZE, 8B39 - PubMed Abstract:
The stabilization of protein-protein interactions (PPIs) has emerged as a promising strategy in chemical biology and drug discovery. The identification of suitable starting points for stabilizing native PPIs and their subsequent elaboration into selective and potent molecular glues lacks structure-guided optimization strategies. We have previously identified a disulfide fragment that stabilized the hub protein 14-3-3σ bound to several of its clients, including ERα and C-RAF. Here, we show the structure-based optimization of the nonselective fragment toward selective and highly potent small-molecule stabilizers of the 14-3-3σ/ERα complex. The more elaborated molecular glues, for example, show no stabilization of 14-3-3σ/C-RAF up to 150 μM compound. Orthogonal biophysical assays, including mass spectrometry and fluorescence anisotropy, were used to establish structure-activity relationships. The binding modes of 37 compounds were elucidated with X-ray crystallography, which further assisted the concomitant structure-guided optimization. By targeting specific amino acids in the 14-3-3σ/ERα interface and locking the conformation with a spirocycle, the optimized covalent stabilizer 181 achieved potency, cooperativity, and selectivity similar to the natural product Fusicoccin-A. This case study showcases the value of addressing the structure, kinetics, and cooperativity for molecular glue development.
- Department of Pharmaceutical Chemistry and Small Molecule Discovery Center (SMDC), University of California, San Francisco, California 94143, United States.
Organizational Affiliation: