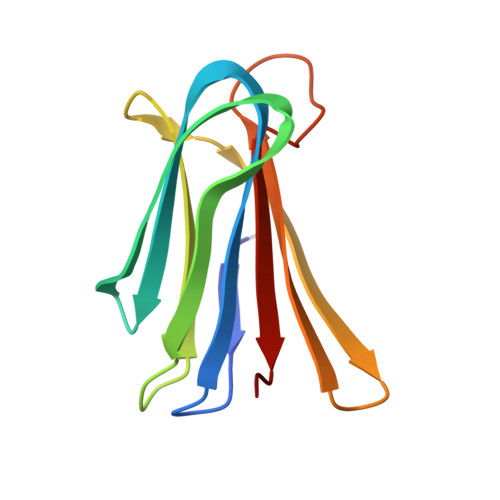
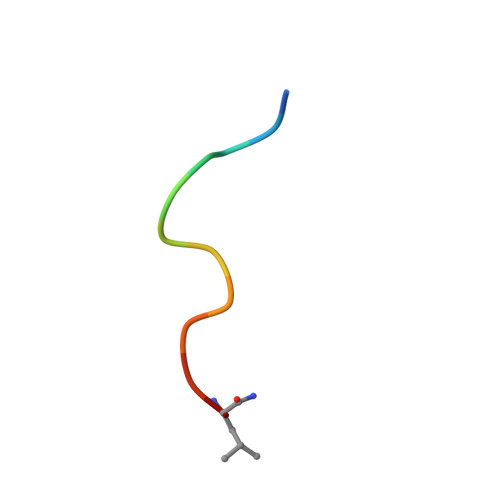
To Fold or Not to Fold: Diastereomeric Optimization of an alpha-Helical Antimicrobial Peptide.
Personne, H., Paschoud, T., Fulgencio, S., Baeriswyl, S., Kohler, T., van Delden, C., Stocker, A., Javor, S., Reymond, J.L.(2023) J Med Chem 66: 7570-7583
- PubMed: 37227046 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.3c00460
- Primary Citation Related Structures:
8AN9, 8ANO, 8ANR, 8AOO - PubMed Abstract:
Membrane disruptive α-helical antimicrobial peptides (AMPs) offer an opportunity to address multidrug resistance; however, most AMPs are toxic and unstable in serum. These limitations can be partly overcome by introducing D-residues, which often confers protease resistance and reduces toxicity without affecting antibacterial activity, presumably due to lowered α-helicity. Here, we investigated 31 diastereomers of the α-helical AMP KKLLKLLKLLL. Three diastereomers containing two, three, and four D-residues showed increased antibacterial effects, comparable hemolysis, reduced toxicity against HEK293 cells, and excellent serum stability, while another diastereomer with four D-residues additionally displayed lower hemolysis. X-ray crystallography confirmed that high or low α-helicity as measured by circular dichroism indicated α-helical or disordered structures independently of the number of chirality switched residues. In contrast to previous reports, α-helicity across diastereomers correlated with both antibacterial activity and hemolysis and revealed a complex relationship between stereochemistry, activity, and toxicity, highlighting the potential of diastereomers for property optimization.
- Department of Chemistry, Biochemistry and Pharmaceutical Sciences, University of Bern, Freiestrasse 3, CH-3012 Bern, Switzerland.
Organizational Affiliation: