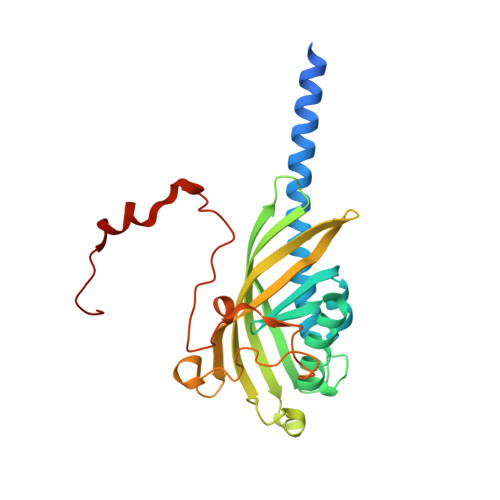
Biochemical and structural characterization of a sphingomonad diarylpropane lyase for cofactorless deformylation.
Kuatsjah, E., Zahn, M., Chen, X., Kato, R., Hinchen, D.J., Konev, M.O., Katahira, R., Orr, C., Wagner, A., Zou, Y., Haugen, S.J., Ramirez, K.J., Michener, J.K., Pickford, A.R., Kamimura, N., Masai, E., Houk, K.N., McGeehan, J.E., Beckham, G.T.(2023) Proc Natl Acad Sci U S A 120: e2212246120-e2212246120
- PubMed: 36652470 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2212246120
- Primary Citation Related Structures:
8ABT, 8ABU, 8ABV, 8ABW - PubMed Abstract:
Lignin valorization is being intensely pursued via tandem catalytic depolymerization and biological funneling to produce single products. In many lignin depolymerization processes, aromatic dimers and oligomers linked by carbon-carbon bonds remain intact, necessitating the development of enzymes capable of cleaving these compounds to monomers. Recently, the catabolism of erythro -1,2-diguaiacylpropane-1,3-diol ( erythro -DGPD), a ring-opened lignin-derived β-1 dimer, was reported in Novosphingobium aromaticivorans . The first enzyme in this pathway, LdpA (formerly LsdE), is a member of the nuclear transport factor 2 (NTF-2)-like structural superfamily that converts erythro -DGPD to lignostilbene through a heretofore unknown mechanism. In this study, we performed biochemical, structural, and mechanistic characterization of the N. aromaticivorans LdpA and another homolog identified in Sphingobium sp. SYK-6, for which activity was confirmed in vivo. For both enzymes, we first demonstrated that formaldehyde is the C 1 reaction product, and we further demonstrated that both enantiomers of erythro -DGPD were transformed simultaneously, suggesting that LdpA, while diastereomerically specific, lacks enantioselectivity. We also show that LdpA is subject to a severe competitive product inhibition by lignostilbene. Three-dimensional structures of LdpA were determined using X-ray crystallography, including substrate-bound complexes, revealing several residues that were shown to be catalytically essential. We used density functional theory to validate a proposed mechanism that proceeds via dehydroxylation and formation of a quinone methide intermediate that serves as an electron sink for the ensuing deformylation. Overall, this study expands the range of chemistry catalyzed by the NTF-2-like protein family to a prevalent lignin dimer through a cofactorless deformylation reaction.
- Renewable Resources and Enabling Sciences Center, National Renewable Energy Laboratory, Golden, CO 80401.
Organizational Affiliation: