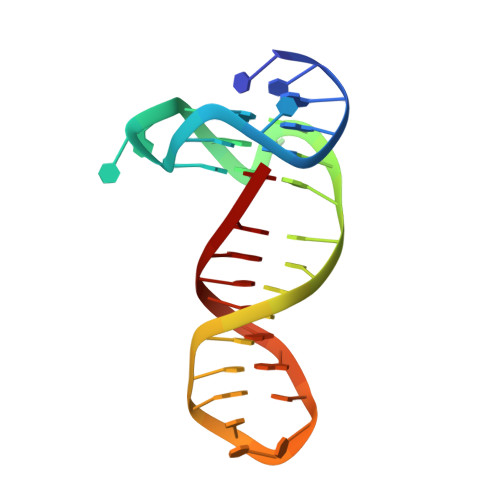
High-affinity binding at quadruplex-duplex junctions: rather the rule than the exception.
Vianney, Y.M., Weisz, K.(2022) Nucleic Acids Res 50: 11948-11964
- PubMed: 36416262 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkac1088
- Primary Citation Related Structures:
8ABD, 8ABN - PubMed Abstract:
Quadruplex-duplex (Q-D) junctions constitute unique structural motifs in genomic sequences. Through comprehensive calorimetric as well as high-resolution NMR structural studies, Q-D junctions with a hairpin-type snapback loop coaxially stacked onto an outer G-tetrad were identified to be most effective binding sites for various polycyclic quadruplex ligands. The Q-D interface is readily recognized by intercalation of the ligand aromatic core structure between G-tetrad and the neighboring base pair. Based on the thermodynamic and structural data, guidelines for the design of ligands with enhanced selectivity towards a Q-D interface emerge. Whereas intercalation at Q-D junctions mostly outcompete stacking at the quadruplex free outer tetrad or intercalation between duplex base pairs to varying degrees, ligand side chains considerably contribute to the selectivity for a Q-D target over other binding sites. In contrast to common perceptions, an appended side chain that additionally interacts within the duplex minor groove may confer only poor selectivity. Rather, the Q-D selectivity is suggested to benefit from an extension of the side chain towards the exposed part of the G-tetrad at the junction. The presented results will support the design of selective high-affinity binding ligands for targeting Q-D interfaces in medicinal but also technological applications.
- Institute of Biochemistry, Universität Greifswald, Felix-Hausdorff-Str. 4, D-17489 Greifswald, Germany.
Organizational Affiliation: