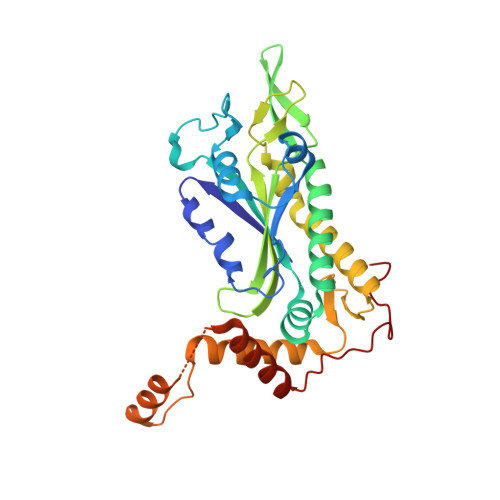
Structural and functional characterization of TraI from pKM101 reveals basis for DNA processing.
Breidenstein, A., Ter Beek, J., Berntsson, R.P.(2023) Life Sci Alliance 6
- PubMed: 36669792
- DOI: https://doi.org/10.26508/lsa.202201775
- Primary Citation Related Structures:
8A1B, 8A1C - PubMed Abstract:
Type 4 secretion systems are large and versatile protein machineries that facilitate the spread of antibiotic resistance and other virulence factors via horizontal gene transfer. Conjugative type 4 secretion systems depend on relaxases to process the DNA in preparation for transport. TraI from the well-studied conjugative plasmid pKM101 is one such relaxase. Here, we report the crystal structure of the trans-esterase domain of TraI in complex with its substrate oriT DNA, highlighting the conserved DNA-binding mechanism of conjugative relaxases. In addition, we present an apo structure of the trans-esterase domain of TraI that includes most of the flexible thumb region. This allows us for the first time to visualize the large conformational change of the thumb subdomain upon DNA binding. We also characterize the DNA binding, nicking, and religation activity of the trans-esterase domain, helicase domain, and full-length TraI. Unlike previous indications in the literature, our results reveal that the TraI trans-esterase domain from pKM101 behaves in a conserved manner with its homologs from the R388 and F plasmids.
- Department of Medical Biochemistry and Biophysics, Umeå University, Umeå, Sweden.
Organizational Affiliation: