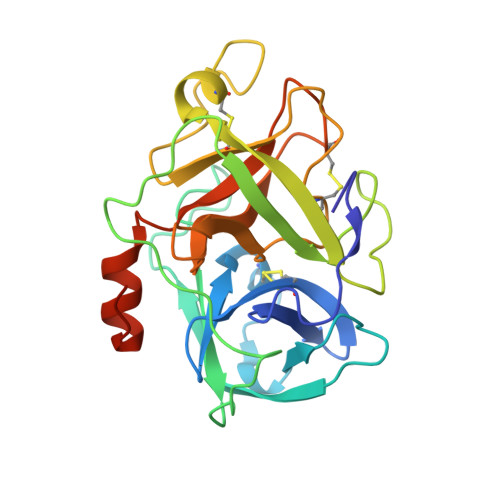
Structural basis of TMPRSS11D specificity and autocleavage activation.
Fraser, B.J., Wilson, R.P., Ferkova, S., Ilyassov, O., Lac, J., Dong, A., Li, Y.Y., Seitova, A., Li, Y., Hejazi, Z., Kenney, T.M.G., Penn, L.Z., Edwards, A., Leduc, R., Boudreault, P.L., Morin, G.B., Benard, F., Arrowsmith, C.H.(2025) Nat Commun 16: 4351-4351
- PubMed: 40348740 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-59677-3
- Primary Citation Related Structures:
8VIS, 9DPF - PubMed Abstract:
Transmembrane Protease, Serine-2 (TMPRSS2) and TMPRSS11D are human proteases that enable SARS-CoV-2 and Influenza A/B virus infections, but their biochemical mechanisms for facilitating viral cell entry remain unclear. We show these proteases spontaneously and efficiently cleave their own zymogen activation motifs, activating their broader protease activity on cellular substrates. We determine TMPRSS11D co-crystal structures with a native and an engineered activation motif, revealing insights into its autocleavage activation and distinct substrate binding cleft features. Leveraging this structural data, we develop nanomolar potency peptidomimetic inhibitors of TMPRSS11D and TMPRSS2. We show that a broad serine protease inhibitor that underwent clinical trials for TMPRSS2-targeted COVID-19 therapy, nafamostat mesylate, was rapidly cleaved by TMPRSS11D and converted to low activity derivatives. In this work, we develop mechanistic insights into human protease viral tropism and highlight both the strengths and limitations of existing human serine protease inhibitors, informing future drug discovery efforts targeting these proteases.
- Structural Genomics Consortium Toronto, Toronto, ON, Canada. Bryanj.fraser@utoronto.ca.
Organizational Affiliation: