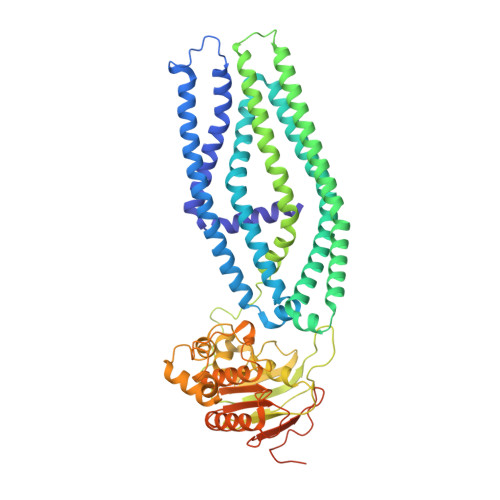
Mechanism of cyclic beta-glucan export by ABC transporter Cgt of Brucella.
Sedzicki, J., Ni, D., Lehmann, F., Wu, N., Zenobi, R., Jung, S., Stahlberg, H., Dehio, C.(2022) Nat Struct Mol Biol 29: 1170-1177
- PubMed: 36456825 Search on PubMed
- DOI: https://doi.org/10.1038/s41594-022-00868-7
- Primary Citation Related Structures:
7ZNU, 7ZO8, 7ZO9, 7ZOA - PubMed Abstract:
Polysaccharides play critical roles in bacteria, including the formation of protective capsules and biofilms and establishing specific host cell interactions. Their transport across membranes is often mediated by ATP-binding cassette (ABC) transporters, which utilize ATP to translocate diverse molecules. Cyclic β-glucans (CβGs) are critical for host interaction of the Rhizobiales, including the zoonotic pathogen Brucella. CβGs are exported into the periplasmic space by the cyclic glucan transporter (Cgt). The interaction of an ABC transporter with a polysaccharide substrate has not been visualized so far. Here we use single-particle cryoelectron microscopy to elucidate the structures of Cgt from Brucella abortus in four conformational states. The substrate-bound structure reveals an unusual binding pocket at the height of the cytoplasmic leaflet, whereas ADP-vanadate models hint at an alternative mechanism of substrate release. Our work provides insights into the translocation of large, heterogeneous substrates and sheds light on protein-polysaccharide interactions in general.
- Biozentrum, University of Basel, Basel, Switzerland.
Organizational Affiliation: