A terminal functionalization strategy reveals unusual binding abilities of anti-thrombin anticoagulant aptamers.
Troisi, R., Riccardi, C., Perez de Carvasal, K., Smietana, M., Morvan, F., Del Vecchio, P., Montesarchio, D., Sica, F.(2022) Mol Ther Nucleic Acids 30: 585-594
- PubMed: 36457701 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.omtn.2022.11.007
- Primary Citation Related Structures:
7ZKL, 7ZKM, 7ZKN, 7ZKO - PubMed Abstract:
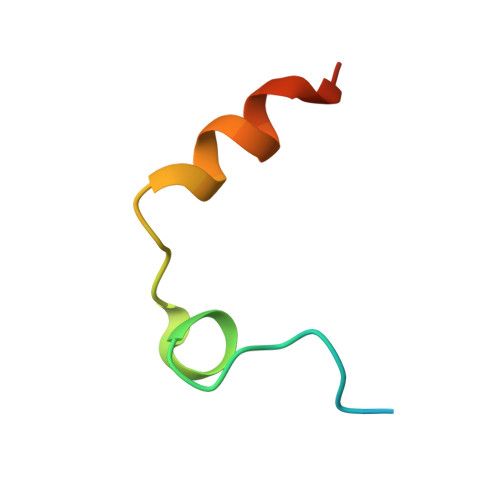
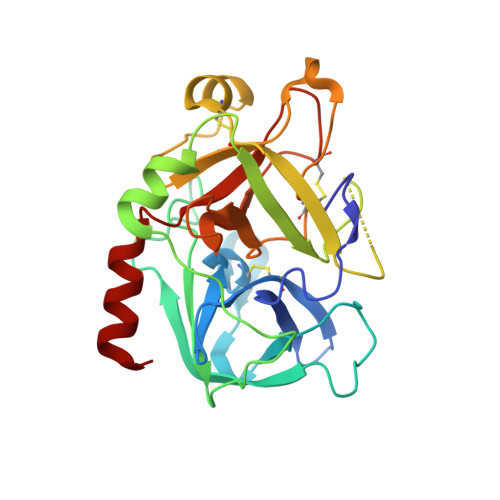
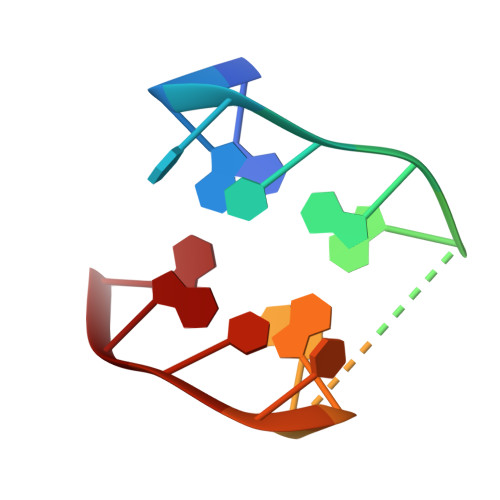
Despite their unquestionable properties, oligonucleotide aptamers display some drawbacks that continue to hinder their applications. Several strategies have been undertaken to overcome these weaknesses, using thrombin binding aptamers as proof-of-concept. In particular, the functionalization of a thrombin exosite I binding aptamer (TBA) with aromatic moieties, e.g., naphthalene dimides (N) and dialkoxynaphthalenes (D), attached at the 5' and 3' ends, respectively, proved to be highly promising. To obtain a molecular view of the effects of these modifications on aptamers, we performed a crystallographic analysis of one of these engineered oligonucleotides (TBA-NNp/DDp) in complex with thrombin. Surprisingly, three of the four examined crystallographic structures are ternary complexes in which thrombin binds a TBA-NNp/DDp molecule at exosite II as well as at exosite I, highlighting the ability of this aptamer, differently from unmodified TBA, to also recognize a localized region of exosite II. This novel ability is strictly related to the solvophobic behavior of the terminal modifications. Studies were also performed in solution to examine the properties of TBA-NNp/DDp in a crystal-free environment. The present results throw new light on the importance of appendages inducing a pseudo -cyclic charge-transfer structure in nucleic acid-based ligands to improve the interactions with proteins, thus considerably widening their potentialities.
- Department of Chemical Sciences, University of Naples Federico II, 80126 Naples, Italy.
Organizational Affiliation: