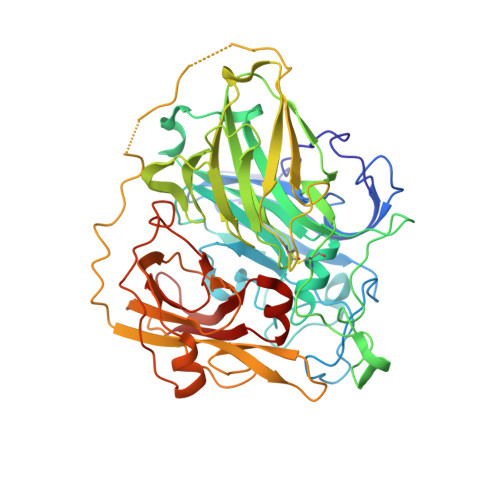
The Structure of Bilirubin Oxidase from Bacillus pumilus Reveals a Unique Disulfide Bond for Site-Specific Direct Electron Transfer.
Gihaz, S., Herzallh, N.S., Cohen, Y., Bachar, O., Fishman, A., Yehezkeli, O.(2022) Biosensors (Basel) 12
- PubMed: 35624560 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/bios12050258
- Primary Citation Related Structures:
7Z5P - PubMed Abstract:
Efficient oxygen-reducing biocatalysts are essential for the development of biofuel cells or photo-bioelectrochemical applications. Bilirubin oxidase (BOD) is a promising biocatalyst for oxygen reduction processes at neutral pH and low overpotentials. BOD has been extensively investigated over the last few decades. While the enzyme's internal electron transfer process and methods to establish electrical communication with electrodes have been elucidated, a crystal structure of BOD from bacterial origin has never been determined. Here we present the first crystal structure of BOD from Bacillus pumilus ( Bp BOD) at 3.5 Å resolution. Overall, Bp BOD shows high homology with the fungal enzymes; however, it holds a unique surface-exposed disulfide bond between Cys229 and Cys322 residues. We present methodologies to orient the T1 site towards the electrode by coupling the reduced disulfide bond with maleimide moiety on the electrodes. The developed configurations were further investigated and revealed improved direct electron transfer rates with the electrodes. The work presented here may contribute to the construction of rationally designed bioanodes or biocathode configurations that are based on redox-active enzymes.
- Department of Biotechnology and Food Engineering, Technion-Israel Institute of Technology, Haifa 3200003, Israel.
Organizational Affiliation: