Targeting non-alcoholic fatty liver disease: Design, X-ray co-crystal structure and synthesis of 'first-in-kind' inhibitors of serine/threonine kinase25.
Kiyeleko, S., Hocine, S., Mautino, G., Kuenemann, M., Nawrotek, A., Miallau, L., Vuillard, L.M., Mirguet, O., Kotschy, A., Hanessian, S.(2022) Bioorg Med Chem Lett 75: 128950-128950
- PubMed: 36030002 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2022.128950
- Primary Citation Related Structures:
7Z4V - PubMed Abstract:
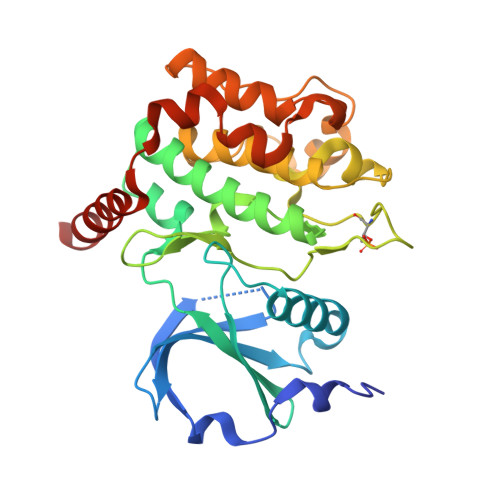
We describe the synthesis of a series of 3-t-butyl 5-aminopyrazole p-substituted arylamides as inhibitors of serine-threonine25 (STK25), an enzyme implicated in the progression of non-alcoholic fatty liver disease (NAFLD). Appending a p-N-pyrrolidinosulphonamide group to the arylamide group led to a 'first-in kind' inhibitor with IC 50 = 228 nM. A co-crystal structure with STK 25 revealed productive interactions which were also reproduced using molecular docking. A new series of triazolo dihydro oxazine carboxamides of 3-t-butyl 5-aminopyrazole was not active against STK25.
- Department of Chemistry, Université de Montréal, Station Centre-Ville, C.P. 6128, Montreal, QC H3C 3J7, Canada.
Organizational Affiliation: