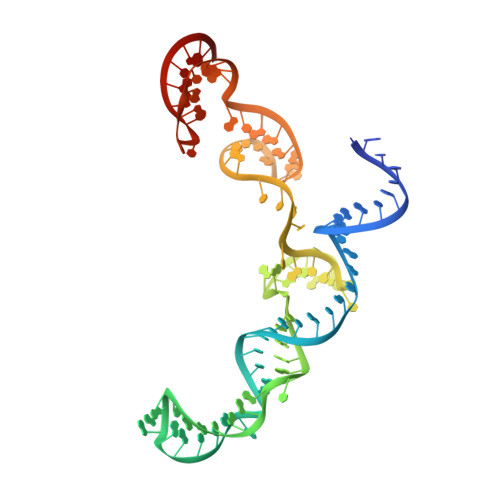
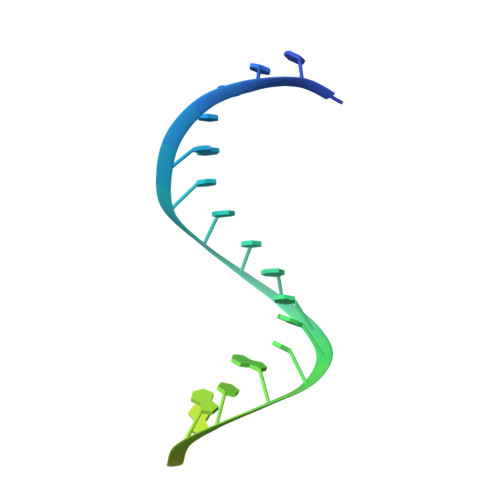
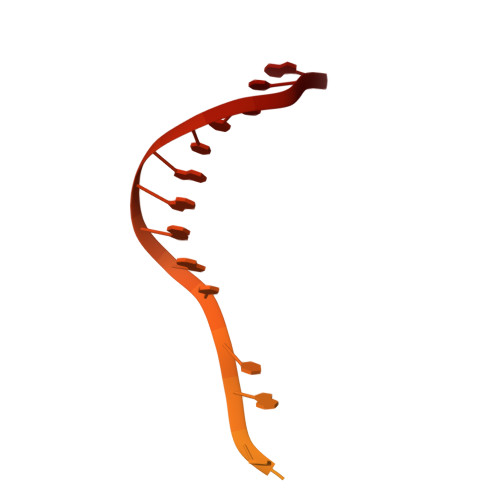
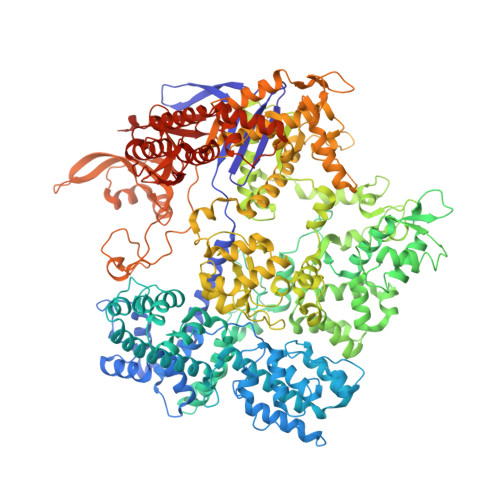
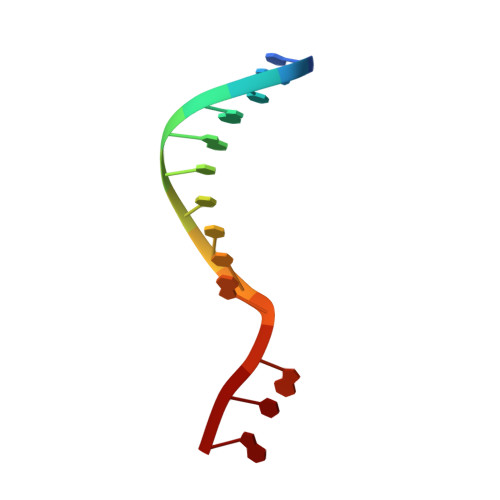
R-loop formation and conformational activation mechanisms of Cas9.
Pacesa, M., Loeff, L., Querques, I., Muckenfuss, L.M., Sawicka, M., Jinek, M.(2022) Nature 609: 191-196
- PubMed: 36002571
- DOI: https://doi.org/10.1038/s41586-022-05114-0
- Primary Citation of Related Structures:
7Z4C, 7Z4D, 7Z4E, 7Z4G, 7Z4H, 7Z4I, 7Z4J, 7Z4K, 7Z4L - PubMed Abstract:
Cas9 is a CRISPR-associated endonuclease capable of RNA-guided, site-specific DNA cleavage 1-3 . The programmable activity of Cas9 has been widely utilized for genome editing applications 4-6 , yet its precise mechanisms of target DNA binding and off-target discrimination remain incompletely understood. Here we report a series of cryo-electron microscopy structures of Streptococcus pyogenes Cas9 capturing the directional process of target DNA hybridization. In the early phase of R-loop formation, the Cas9 REC2 and REC3 domains form a positively charged cleft that accommodates the distal end of the target DNA duplex. Guide-target hybridization past the seed region induces rearrangements of the REC2 and REC3 domains and relocation of the HNH nuclease domain to assume a catalytically incompetent checkpoint conformation. Completion of the guide-target heteroduplex triggers conformational activation of the HNH nuclease domain, enabled by distortion of the guide-target heteroduplex, and complementary REC2 and REC3 domain rearrangements. Together, these results establish a structural framework for target DNA-dependent activation of Cas9 that sheds light on its conformational checkpoint mechanism and may facilitate the development of novel Cas9 variants and guide RNA designs with enhanced specificity and activity.
- Department of Biochemistry, University of Zurich, Zurich, Switzerland.
Organizational Affiliation: