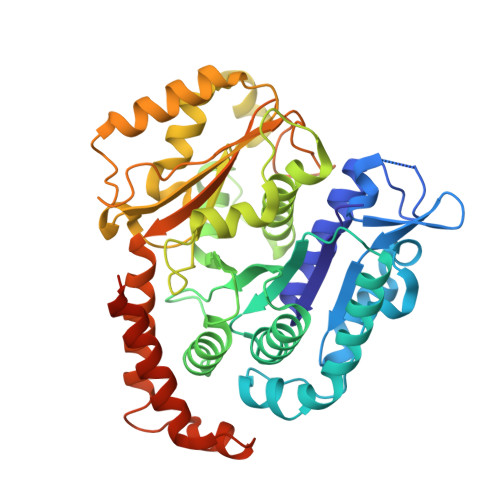
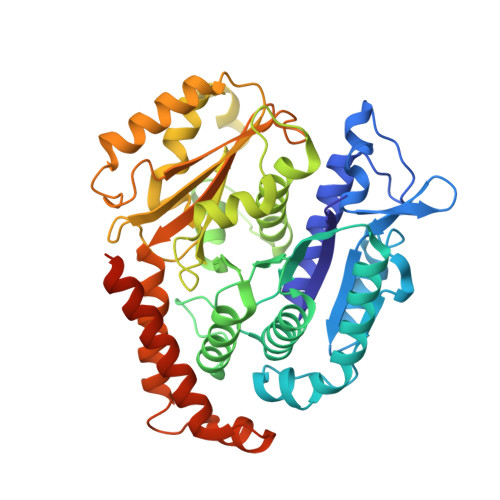
Structural insights into the mechanism of GTP initiation of microtubule assembly.
Zhou, J., Wang, A., Song, Y., Liu, N., Wang, J., Li, Y., Liang, X., Li, G., Chu, H., Wang, H.W.(2023) Nat Commun 14: 5980-5980
- PubMed: 37749104 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-41615-w
- Primary Citation Related Structures:
7YSN, 7YSO, 7YSP, 7YSQ, 7YSR - PubMed Abstract:
In eukaryotes, the dynamic assembly of microtubules (MT) plays an important role in numerous cellular processes. The underlying mechanism of GTP triggering MT assembly is still unknown. Here, we present cryo-EM structures of tubulin heterodimer at their GTP- and GDP-bound states, intermediate assembly states of GTP-tubulin, and final assembly stages of MT. Both GTP- and GDP-tubulin heterodimers adopt similar curved conformations with subtle flexibility differences. In head-to-tail oligomers of tubulin heterodimers, the inter-dimer interface of GDP-tubulin exhibits greater flexibility, particularly in tangential bending. Cryo-EM of the intermediate assembly states reveals two types of tubulin lateral contacts, "Tube-bond" and "MT-bond". Further, molecular dynamics (MD) simulations show that GTP triggers lateral contact formation in MT assembly in multiple sequential steps, gradually straightening the curved tubulin heterodimers. Therefore, we propose a flexible model of GTP-initiated MT assembly, including the formation of longitudinal and lateral contacts, to explain the nucleation and assembly of MT.
- State Key Laboratory of Membrane Biology, Tsinghua University, Beijing, 100084, China.
Organizational Affiliation: